Research Article
Potential of Developed Microbial Biofilms in Generating Bioactive Compounds
Institute of Fundamental Studies, Hantana Road, Kandy, Sri Lanka
G. Seneviratne
Institute of Fundamental Studies, Hantana Road, Kandy, Sri Lanka
The microbial world has many diverse relationships that play important roles in human lives. Microbially-produced bioactive compounds are attracting increased attention as useful agents for medicine, veterinary medicine, agriculture and as unique biochemical tools and have been isolated using both mono and mixed cultures of microbes (Borowitzka, 1995; Omura, 1992; Sankpal and Kulkarni, 2002).
There is a great scope for developing a biofilm technology of producing eco-friendly, beneficial microbial biofilms (EBM) for numerous applications (Seneviratne, 2003), because microbes in their ‘biofilm’ mode have already been shown to efficiently perform various biotransformation processes (Seneviratne and Jayasinghearachchi, 2005; Seneviratne et al., 2006). Biofilms are multi-cellular consortia of microbial cells (fungal, bacterial, algal and/or other microbial), which closely adhere to each other and are very often encased in a self-produced Extra-cellular Polymeric Substance (EPS). Such interactions give the biofilm community metabolic and physiological capabilities which are not possible for the individual unattached cells. Although biofilms are generally considered to be problematic, their beneficial aspects have often been overlooked.
Previous studies show that the production of organic acids was increased in biofilms compared to monocultures and non-biofilm forming mixed cultures (Bandara et al., 2006). Biofilms have a unique pattern of gene expression, which is different from their non-biofilm forming stages (Vilain and Brözel, 2006). Thus, it is apparent that biofilms have a higher potential of producing new organic acids, some of which may be bioactive compounds.
There is limited knowledge of interspecies interactions in biofilm communities (Cowan et al., 2000). Yet, a range of metabolic interactions has been observed among microorganisms in biofilms, including mutualistic and commensal relationships (Møller et al., 1998; Wolfaardt et al., 1994). An understanding of these relationships in biofilms could aid the design of multispecies biofilms for the biosynthesis of specialty chemicals or the biodegradation of xenobiotics that cannot be metabolized efficiently by a single microorganism (Seneviratne et al., 2006; Cowan et al., 2000).
Therefore, in this study we attempt to use the biofilm technology to produce bioactive compounds by using a previously developed fungal-bacterial biofilm (Seneviratne and Jayasinghearachchi, 2005). Monosaccharides constitute a major group of bioactive compounds, because they are the starting materials for a number of drugs such as those possessing anti-microbial, anti-tumor, anti-viral, anti-oxidant, free radical scavenging and anti-inflammatory activities (Methacanon et al., 2005). Thus, monosaccharides are used as the test compounds in the present study.
Culturing of Microorganisms
Bradyrhizobium elkanii SEMIA 5019 applied as a soybean inoculant was used for this study because microbial metabolism in nature is frequently nitrogen limiting and this N2 fixer has been shown to accumulate atmospheric N2 in microbial communities (Jayasinghearachchi and Seneviratne, 2004a, b). Cultures were maintained on Yeast Mannitol Broth (YMB) (Somasegaran and Hoben, 1994), but without agar. They were incubated at 28°C for 6 days on a rotary shaker. Penicillium sp. isolated from garden soil (Seneviratne and Jayasinghearachchi, 2005) was used as the fungal counterpart of the biofilm and the fungal cultures were maintained on Saboraud’s agar.
Preparation of Treatments
One milliliter of a 6 day old B. elkanii SEMIA 5019 culture was inoculated together with 50 μL Penicillium sp. spore suspension to 50 mL of concentrated YMB (Jayasinghearachchi and Seneviratne, 2004a) to form a mixed culture. They were separately inoculated to 50 mL of the concentrated YMB to form monocultures. Fifty milliliter of the concentrated YMB without microorganisms was used as the control. Each treatment was replicated four times in 50 mL conical flasks and kept on a rotary shaker at 28°C for 14 days. Biofilms were observed after 7 days of incubation using a phase-contrast microscope with an oil immersion lens and lacto-phenol cotton blue stain. After 14 days of incubation, the cultures were centrifuged at 13,148x g for 20 min and supernatants were separated and freeze-dried.
Sample Preparation
Analysis of monosaccharides was done by gas-liquid chromatography of alditol acetates based on the method of Amelung et al. (1996). In this procedure, carbohydrates with acid constituents are hydrolysed to constituent monosaccharides, which are subjected to reduction followed by acetylation to yield alditol acetates. Of the freeze-dried crude material, 0.2 g was hydrolyzed with 0.6 mL of 4M trifluoroacetic acid (TFA) at 105°C for 4 h. It was rested overnight and the hydrolysate was filtered through a glass fibre filter. It was evaporated to dryness by a stream of N2 at 50°C, to remove TFA. One milliliter of methanol was added and evaporated to dryness and this was repeated. Resulting monosaccharides were reduced to alditols using 1M NaBH4 in NH4OH and kept at 25°C for 30 min. To the solution, 0.3 mL acetic acid in methanol was added to decompose excess NaBH4 until bubble formation stopped. The solution was blown to dryness by a flow of N2 and above step was repeated. Borohydrate was removed by the addition of 0.5 mL of methanol and dried under a stream of N2. To the reduced samples, 10 μL of acetic anhydride and 100 μL of pyridine were added and reacted at 100 °C for 20 min to complete acetylation. After that, 0.5 mL of toluene was added and blown to dryness with a stream of N2. This was repeated. To this, 0.5 mL distilled water and 0.5 mL ethyl acetate were added and mixed by turning up and down 3 times. The upper layer was removed using a Pasteur pipette and filtered through a silica gel filter. This was re-extracted with 0.5 mL ethyl acetate and concentrated in a Nunc cryo tube.
Gas-liquid Chromatographic Analysis
Gas-liquid chromatography of the monosaccharides was performed with a Shimadzu GC-9 AM Gas Chromatograph (GC) with a 30x0.25 mm i.d. x 0.25 μm film thickness SPB™ -1701 fused silica capillary column. The analytical conditions were an initial temperature of 180°C, injector at 220°C and the detector at 260°C. Residue obtained from the sample preparation was dissolved in 30 μL ethyl acetate of which 20 μL was injected into the GC by the split technique. Standards of monosaccharides were also analyzed with similar preparation steps but without TFA hydrolysis.
Mycelium of the Penicillium sp. was heavily colonized by B. elkanii SEMIA 5019, forming the Penicillium sp.-B. elkanii SEMIA 5019 biofilm (Fig. 1). Detectable quantities of the monosaccharides fucose, ribose, arabinose and xylose were produced only by the biofilm (Table 1). They were not detected in the B. elkanii SEMIA 5019 alone or Penicillium sp. alone treatments, possibly due to their production below the detection limits. The monocultures produced up to 10 detectable monosaccharides, whereas the biofilm produced 123 of them, though most of them are yet to be identified. This clearly shows the very high potential of the biofilm in generating bioactive compounds. The variabilities of the types and quantities of the compounds among replicates of the biofilm treatment, as reflected from the standard errors (Table 1) could be attributed to the method of fungal inoculation, because the fungus carries variable numbers of genetically different spores. This could be overcome by inoculating a piece of mycelium grown from a single spore of the fungus.
Previous studies show that the expressions of polysaccharide-producing genes are upregulated in biofilm cells compared with planktonic (freely swimming single) cells in liquid media (Davies et al., 1993). The ability of biofilms to produce higher detectable amounts of monosaccharides in comparison to monocultures may be rendered by a change in gene expression of individual cells when in the biofilm relationship.
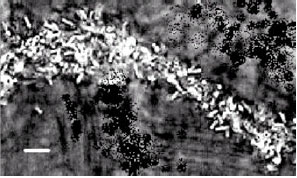 | |
| Fig. 1: | Phase-contrast microscopic view of Penicillium sp.-Bradyrhizobium elkanii SEMIA 5019 biofilm. A mycelial filament of the Penicillium sp. has been heavily colonized by B. elkanii SEMIA 5019. Horizontal bar is 3 μm |
| Table 1: | Concentrations of monosaccharides detected of four replicates of the microbial treatments |
 | |
| Mean±standard error. ND: Not Detected. Ranges are the numbers of the four replicates | |
In the biofilm there is a range of conditions surrounding the cells due to heterogeneity of the consortium structure (Christensen et al., 2002). Metabolic interactions between community members occur in all regions of the biofilm independent of the local architecture (Møller et al., 1998). Non-biofilm forming mixed culture systems require active motility and cell-to-cell communication to achieve optimal conditions. Thus, substrates are more optimally utilized in the biofilm formation, resulting in faster degradation of the primary nutrients (Christensen et al., 2002). This is further enhanced, by the critical cell density dependent quorum sensing that leads to biofilm formation (Kong et al., 2006). In previous studies, single species biofilms have been used to produce industrially important chemicals (Qureshi et al., 2005). However, the use of dual-species biofilms have higher quorum sensing and community level gene expression that lead to higher levels of substrate utilization and maximization of products (Cowan et al., 2000), as was seen in the present study.
Conventional drug discovery studies use natural or genetically modified monocultures of microorganisms. But in the biofilm technology interactions of the same microorganisms with different microbes can yield a multitude of results and an endless number of novel compounds. This technology can be used to produce different compounds by the combination of a variety of microbes, by changing the carbon source or altering the culture conditions. Studying the concerned interactions and their culture conditions can also optimize individual compounds with bioactivity. Its manipulative ability therefore can be made use to open a myriad of avenues for the discovery of novel compounds that can be used in drug discovery studies, thus finding solutions to many hurdles in the treatment of diseases. The advantage of this method also lies in the reduced time frame compared to the isolation of compounds from plants and conventional methods used for these isolations. Further studies to optimize this microbial technology and also to produce and detect other compounds of bioactive significance are in progress.
Microbial studies of the project were initiated during the Sri Lanka-Belgium collaboration on biological nitrogen fixation (1991-1997). We thank Ms. R. Ratnayake for her guidance on GLC. The assistance rendered by Ms. K. Karunaratne, Anjani Weerasekara and Mr. Manoj Bandara, K. Karunadasa is much appreciated.