Research Article
Isolation and Characterization of 2,4-Dichlorophenoxyacetic Acid Degrading Organisms from Soil in Jordan Valley
Department of Biotechnology, Faculty of Agriculture Technology, Al-Balqa Applied University, Jordan
Large amounts of man-made chlorinated organic chemicals have been used in agriculture as herbicides and pesticides. Among these, 2-4-dichlorophenoxyacetic acid (2,4-D) has received widespread use as a herbicide for more than 40 years.Unlike many of the synthetic compounds released into the environment, 2,4-D is rapidly mineralized by soil bacteria (Biederbeck et al., 1987), (Foster and Mckerches, 1973).The sizes of populations of microorganisms able to degrade 2,4-D have been estimated by the most-probable-number method (Fournier, 1980); (Sandman and Loos, 1984). Organisms that have been reported to be capable of degrading 2,4-D belong to a number of genera, including Achromobacter, Alcaligenes, Arthrobacter, corynebacterium, Flavobacterium, Pseudomonas and Streptomyces (Bell, 1957; Don et al., 1985; Loos et al., 1967; Walker and Newman, 1956).
Xenobiotics have been used extensively during the past 5 decades both in industry and agriculture. This use has led to the accumulation of these compounds in the environment polluting natural resources (Harker et al., 1989; Kukor et al., 1989).The widespread use of pesticides in agriculture to increase crop production has generated a series of toxicological and environment problems. The use of herbicides has increased in the last few years to replace mechanical weeding (Miwa and Kuwatsuka, 1991). Chlorinated phenoxy acid derivatives (esters and amines) are the most common and the most widly used herbicides 2,4- Dichlorophenoxyacetic acid (2,4- D) is one that was used to defoliate jungle areas in South Vietnam as a component of the "Orange Agent" (Oliveira and Palermo-Neto, 1995).
On the other hand, pollution has induced rapid evolution of specific biodegradative pathways in terrestrial and aquatic bacteria. Several bacterial genera can degrade and metabolize 2,4-D, Alcaligenes, Pseudomonas, Acenitobacter, Arthrobacter and Corynebacterium (Don and Pemberton, 1981; Don and Pemberton, 1985; Streber et al., 1987; Suwa et al., 1996). However, abiotic factors such as phtooxidation and adsorption of 2,4-D to particulates could also contribute to the degradation of some toxic substances partially but never to mineralization (Amy et al., 1985).
It was reported that degradation of 2,4-D was due to plasmid carried genes in the strains of Pseudomonas and Alcaligenes, furthermore, these plasmids often carry genes responsible for degradation of other structurally related chlorinated organics (Don and Pemberton, 1985; Don et al., 1985; Greer et al., 1990).
The main objective of this study was to isolate indigenous strains of bacteria that are capable of degrading certain local pollutants. This is due to the fact that these strains possess the appropriate metabolic, physiological, and kinetic characteristics that are highly adapted to the existing environmental conditions under which bioremediation takes place (Greer et al., 1990; Greer and Shelton, 1992). If such indigenous organisms were not obtained, microbes containing the necessary genes are introduced into the polluted environment in a process called bioaugmentation. A such process faces two problems: first the introdused organism may die in few weeks due to boitic and abiotic stresses. Secondly, the introduced organism may not be able to spread easily through the soil medium. The transfer of genes responsible for degradation to indigenous soil recipients may help to overcome such difficulties (Digiovanni et al., 1996).
Although the import of herbicides and pesticides in Jordan is controlled and restricted, the use of these chemicals is still far from being regulated. This would eventually lead to the pollution of the limited natural resources (Ministry of Planning, 1994). Since the environmental conditions in Jordan - especially in the Jordan Rift Valley – are unique (the lowest point on Earth), this may contribute largely to the existence of unique and undiscovered strains of bacteria that are able to degrade 2,4-D and other pesticides. These strains could have potent degradative capabilities. For these reasons, the presence of such strains in the Jordanian environment was investigated.
Media preparation
Minimal salt medium (MSM) was used throughout this study (Kelly et al., 1989). The contents are grams per liter of distilled H2O:2,4-D (0.25), K2HPO4 (0.5), (NH4)2 SO4(0,5), MgSO4.7H2O (0.5), FeCl3.6H2O (0.01), CaCL2.2H2O (0.01), MnCl2.4H2O (0.01) and ZnSO4.7H2O (0.0001).The medium was boiled for 10 minutes and autoclaved at 15 bars and 1210C without significant loss in 2,4-D (final pH was 6.95) (Cullimore, 1981).Nutrient agar medium containing 250 ppm 2,4-D was prepared. The media used in characterization of isolated strains of 2,4-D degraders were prepared as described in Collins and Lyne’s Microbiological Methods (Collins et al., 1995).
Processing of soil samples
Soil samples from the upper 10 cm of soil were taken from the different sites and kept in plastic bags at ambient temperature until processing (Cullimore,1981; Ka et al., 1994). The bags were kept open to prevent the increase of their temperature and subsequently death of bacteria. Twenty grams of thoroughly mixed soil sample from each site, was homogenized in 200 ml sterile saline solution (0.85%) by shaking the preparation on a water bath shaker (Model G76, New Brunswick Scientific Co., Edison, N.J., USA) at 200 rpm and room temperature for 20 min. (Ka et al., 1994).
Enumeration of 2,4- D degraders
To carry out most-probable-number (MPN) analysis for estimating 2,4–D degraders in soil samples, the following steps- which were obtained from (Eaton et al., 1995) with some modifications-were performed and they were repeated for each site:
| 1) | The medium used in MPN analysis was MSM containing filter sterilized 0.1 μg cycloheximide/ml to prevent fungal growth (Top et al., 1995). |
| 2) | Each site had 15 screw capped tubes;10 tubes contained 3 ml of MSM supplemented with 250ppm 2,4-D (single strength) and the other 5 contained also 3 ml of MSM but were supplemented with 500 ppm 2,4-D (double strength). |
| 3) | Five of the single strength tubes were inoculated with 0.1 ml of soil suspension and the other 5 tubes were inoculated with 1.0 ml of soil suspension from the same site. |
| 4) | The double strength tubes were inoculated with 10.0 ml of soil suspension. |
| 5) | The inoculated tubes were incubated at 30°C and shaken at 150 rpm in a controlled environment incubater shaker (New Brunswick Scientific Co., N.J., USA) for 6 weeks in the dark (Amy et al., 1990; Digiovanni et al., 1996; Lechmicke et al., 1979). |
| 6) | The inoculated tubes were tightly closed to prevent excessive dehydration, but were loosened every 3-4 days to allow gaseous exchange (Cullimore, 1981). |
| 7) | At the end of the incubation period, duplicates of 1.5 ml of the MPN medium from each tube were transferred into 2 Eppendorf tubes centrifuged for 15 min in a microcentrifuge (Sigma, Model 3K30, Sigma Laboratory Centrifuges GmbH, Germany) at 16,000xg and 4°C (Digivanni et al., 1996). |
| 8) | High-performance liquid chromatography (HPLC) was performed on the supernatants with positive tubes being scored as those exhibiting more than 25% decrease in the initial concentration (Ci) of 2,4-D (Cullimore, 1981). |
| 9) | The population density of 2,4-D-degraders was calculated from MPN tables provided by (Collins et al., 1995). |
Isolation of 2,4-D-degraders
For each site, a 100 ml screw capped flask containing 67.5 ml of MSM, 250 ppm 2,4-D and 0.1 μg/ml filter sterilized cycloheximide was prepared in addition to the sets of MPN tubes as control (without 2,4-D). A volume of soil suspension (7.5 ml) was transferred to each flask and incubated together with the MPN tubes. After 6 weeks, 7.5 ml from each flask were transferred into 67.5 ml of freshly prepared MSM containing 250 ppm 2,4-D and was re-incubated at the same conditions for 7 days. The enriched cultures were streaked onto 2,4-D agar medium plates and incubated at 30°C. Mixed cultures were purified on 2,4-D agar medium and pure colonies were preserved by freezing at –20°C in sterile 25% glycerol.
Characterization of bacterial isolates
The isolated strains were characterized using standard procedures described in Collins and Lyne’s Microbiological Methods (Collins et al., 1995) and Bergy’s Manual of Systematic Bacteriology (Palleroni, 1984).The oxidase test was performed by placing a small quantity of bacteria on a filter paper soaked with 1% tetramethyl-p-phenylenediamine dihydrochloride dye. This dye (colorless in its reduced state) is rapidly oxidized to a violet or purple colour by oxidase- positive species that contain c-type cytochrome. Biochemical tests were performed using standard methods as well as commercial rapid test system, NEFERM test (Lachema, Czech Republic), which is intended for the diagnostics of Gram-negative non-fermenting bacteria, including Pseudomonas, Acenitobacter, Xanthomonas, Flavobacterium and others. NEFERM test is based on the principle of dehydrated diagnostic media that are deposited in microwells on a microtitration plate (each plate is enough for 8 isolates). On each plate there are 12 fundamental teste: arginine and esculine hydrolysis, indole formation, citrate utilization, phosphatase and urease production, glucose, mannitol, xylose, maltose and lactose assimilation and nitrite reduction. Complete identification by means of NEFERM test was by carrying out the test for oxidase and the beta-galactosidase test. After 24 h of incubation at 30°C, identification of the unknown organism is made by using the differential tables and the identification code leaflet supplied by the manufacturer.
Selecting the most potent 2,4-D-degrader
All the isolates were first enriched in screw capped tubes containing 10 ml nutrient broth supplemented with 250 ppm 2,4-D and incubated at 30°C and shaken at 150 rpm overnight. Seven point five ml of enriched cultures were transferred into 100 ml flasks containing 67.5 ml MSM and 250 and 500 ppm 2,4-D. The inoculated flasks were incubated at 30°C and 150 rpm. Duplicates of 1.5 ml from each flask were withdrawn aseptically every 24 h into Eppendorf tubes, then centrifuged and analyzed by HPLC for 2,4-D degradation.
High-performance liquid chromatograpy (HPLC) analysis
Twenty microliters aliquates from culture supernatants, were analyzed with an HP1100 liquid chromatograph (Hewlett-Packard Co., Palo Alto, California). The column used was an RP-18 3.0 ml H3PO4 per liter distilled water (pH 2.5). The mobile phase was filtered with a 0.45 μm-pore- size filter and was degassed with helium prior to use. Under these conditions the HPLC system had a detection limit of 1 ppm for 2,4-D. Concentrations were dedused from areas of sample peaks compared with those of known standards prepared in 0.1M phosphate buffer (pH 7.0) and similarly run (Haugland et al., 1990; Kelly et al., 1989).
Morphological and physiological characteristics of isolated strains
Seven bacterial isolates were the outcome of purification of mixed cultures from all the sites. The pure culture of one isolate from site 1 was found to degrade initial concentration of 2,4-D completely after 45 h of incubation. The 2,4-D degrader was identified as Pseudomonas putida.
This isolate was identified and characterized according to Microbiological methods (Collins et al., 1995) and Bergy’s Manual of systematic Bacteriology (Palleroni, 1984).
Results of biochemical tests for Psuedomonas putida shown in Table 1. NEFERM test, which is intented for the diagnostic of gram negative non-ferminting bacteria, was conducted for identification of the 2,4-D degrader.
MPN enumeration of 2,4-D-degraders
The numbers of 2,4-D-degrading organisms, in both the exposed and the no-history soils, exhibited a large variation. Site 1, which had a previous history of exposure to 2,4-D, had 88 2,4-D- degraders/g soil. Sites 2,3,4 and 5 showed very low numbers (Table 2).
On the other hand, the flasks that were incubated with the MPN tubes inoculated with the soil suspension from each site became turbid even though they did not show any significant decrease in the initial concentration of 2,4-D except for site 1 which depleted 2,4-D entirely within 45 h of incubation.
Biodegradation of 2,4-D (HPLC analysis)
The characterized 2,4-D degrader (Pseudomonas putida) showed that it has ability to metabolize and minerelize 500 ppm of 2,4-D in 45 h when incubated at 30°C (Fig. 1).
When low levels (200 ppm) of 2,4-D were used in enrichment cultures it was totally disappeared in 24 h (Fig. 2).
Toxicity of 2,4-D at higher concentration (1000ppm) was observed. The concentration of 2,4- D did not change (1000ppm) (Fig. 3) and colonies of 2,4-D degradores were not detected.
In this study we succeeded in isolating 2,4-D metabolizing organism (Pseudomonas putida) from Jordan Valley soil. The strategy of using progressively less organic supplement and higher concentration of 2,4-D in each of the enrichment steps seems to have worked, although it should be noted that it did not work in all cases.
| Table 1: | Biochemical reactions of non-fermentative Gram-negative soil isolates | ||
 | |||
| aReactions obtained using NEFERM test plates after 24 h incubation at 30°C bRefer to Materials and Methods and Collins et al. (1995) | |||
| Note: | + | = | 86 to 100% positive reactions |
| (d) | = | 25 to 75% positive reactions | |
| - | = | 0 to 14% positive reactions | |
| Table 2: | MPN of 2,4-D-degraders in the sites studied (the numbers of positive tubes were deduced according to (Eaton et al., 1995) |
 | |
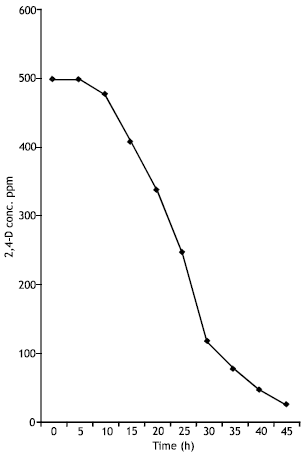 | |
| Fig. 1: | Biodegradation of 500 ppm of 2,4-D by Pseudomonas putida |
One of the most intresting features of this isolate is the fact that it is slow-growing microorganism and sensitive to high concentrations of nutrients.
Itappears that when undiluted environment sample are enriched in both cultures containing a higfh 2,4-D concentration, only fast growing degraders outgrow slow growers in the cultures (Ka et al., 1994).
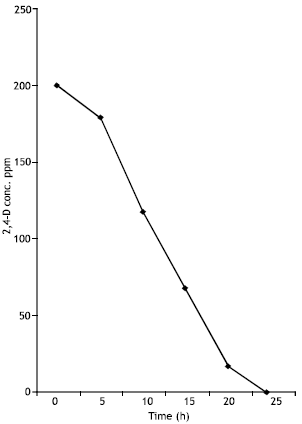 | |
| Fig. 2: | Biodegradation of 200 ppm of 2,4-D by Pseudomonas putida |
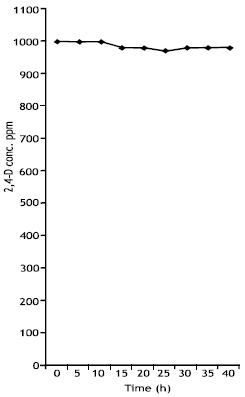 | |
| Fig. 3: | Toxic effect of 1000 ppm of 2,4-D on Pseudomonas putida |
Since only slow growers were obtained from our enrichment cultures without diluting soil samples, there are apparently no fast-growing 2,4-D degrading microorganisms in these soils.
Growth of 2,4-D-degraders
The relatively high number encountered at site 1 is very logical; since this site is the only one that had been exposed to 2,4-D as reported previously (Ministry of Planning, 1994). This finding was supported by several previous studies (Holben et al., 1992) observed a sharp increase in 2,4-D-metabolizing populations in response to amendment of different types of soils with 2,4-D in the laboratory. It was reported that the frequent exposure of soils to 2,4-D would appear to enhance the degrader population and the non-treated soils contained fewer degrading organisms than treated ones.
The relatively longer period for complete degradation of 2,4-D that was observed in the flask containing the mixed culture of site 1 in relation to the pure isolate from the same site – which degraded 2,4-D completely in 45 h – was also noticed by Lewis et al., 1984). He explained that the presence of various cultures, filtrates, or exudates of algae, fungi, or other bacteria can either stimulate or inhibit bacterial transformation of 2,4-D. In this study, there was a delayed onset of 2,4-D biodegradation. Nevertheless, we cannot generalize since the biodegradation activity in other sites was limited.
The fact that other flasks – containing either mixed or pure cultures – showed growth in the culture medium without degradation of 2,4-D, during the of period study consisted with the finding of Ka et al. (1994). He reported the isolation of bacterial isolates from fields treated with low concentration of 2,4-D showed slow growth and could not be recovered from fields treated with high concentration of 2,4-D; some of these active cultures failed to degrade 2,4-D and others degraded 2,4-D slowly in relatively longer periods of incubation (Ka et al., 1994).
Identification of bacterial isolates
Characterization studies of the isolates from the experiments in this work, as well as those by other reserchers (Kilpi, 1980; Lappin et al., 1985; Smith-Grenier and Adkins, 1996), indicated that Pseudomonas species and Pseudomonas – like species of the genera Sphingomonas, Chryseomonas and Xanthomonas consistently made up significant proportion of phenoxy acid herbicide-degrading communities. Mixed cultures capable of mineralizing 2,4-D, mecocorp, or dichloroprop contained either fluorescent pseudomonas (Kilpi, 1980) or 60% Pseudomonas species and 25% Xanthomonas maltophilia (Lappin et al., 1985).
Pseudomonads are a vast heterogeneous group of bacteria that occur in substantial numbers in the soil where they are active agents of minerlization of organic matter. Most species grow well in simple minimal medium with a single organic compound as carbon and energy source (Palleroni, 1992). Pseudomonads possess a variety of diverse catabolic pathways that enable them to metabolize an equally diverse number of low molecular weight organic substrates, including chlorinated aliphatic hydrocarbons such as the phenoxyalkanoic acid herbicides (Smith-Grenier and Adkins, 1996). Some species of pseudomonads, notably P. cepacia, are capable of using 100 different organic compound for growth (Kilbane et al., 1982).
The presence of substantial numbers of nutritionally versatile pseudomonads in soil likely accounts for the reason why these microorganisms are so readialy isolated from soil. Furthermore, experimental conditions of standard enrichment culture techniques, as those of this research, are no doubt optimal for the selection of pseudomonads and related species. It is possible that under a different set of enrichment procedures, a different group of microorganisms might have been isolated (Amy et al., 1985; Evans et al., 1971; Fisher et al., 1978; Greer and Shelton, 1992; Ka et al., 1994; Miwa and Kuwatsuka, 1991).
Biodegradation of 2,4-D
Pseudomonas putida showed it ability to entirely degrade 2,4-D at concentration below 500 ppm, but higher concentration (1000 ppm) become toxic and inhibit the growth of the bacteria.
Our research indicates several important points. First, 2,4-D degrading microorganisms were widespread in soil environment but were difficult to culture (Fulthrope et al., 1996). Second, most of 2,4-D degraders were slow-growing microorganisms and were sensitive to high concentrations of organic nutrients. Third, high concentration of 2,4-D (1000 ppm) was very toxic for Pseudomonas putida. Fourth, the isolation of 2,4-D degradors from these soils might be suggested that selection for chloroaromatic degradation has occurred. Further work will focus on the metabolic pathway of 2,4-D degradation and genetic analysis of 2,4-D degrading gene (s) in Pseudomonas putida which we have isolated.
This reserch was supported by the Faculty of Graduate Studies and Scientific research at Al-Balq’a Applied University.