Research Article
Fungally-transmitted Rod-shaped Plant Viruses: Biology, Transmission and Molecular Pathology
Department of Plant Pathology, Faculty of Crop Protection Sciences, N.W.F.P. Agricultural University, Peshawar, Pakistan
The genus name, furovirus, was approved by International Committee on the Taxonomy of viruses (ICTV) for fungally-transmited, rod-shaped plant viruses. All furoviruses possess a divided genome composed of two or more components of single-stranded (ss) RNA and are transmitted in moist soil by motile zoospores of plasmodiophorid 'fungus'. The occurrence of the viruses are worldwide particularly in temperate regions (Table 1). The natural vectors of these viruses belong to 2 genera Polymyxa and Spongospora. Soil-borne wheat mosaic virus (SBWMV) is the type species of the furovirus genus, which include other species such as oat golden stripe virus (OGSV), peanut clump virus (PCV), potato mop-top virus (PMTV), sorghum chlorotic spot virus (SgCSV). Beet necrotic yellow vein virus (BNYVV), beet soil-borne virus (BSBV), broad bean necrosis virus (BBNV), Hypochoeris mosaic virus (HyMV) and rice stripe necrosis virus (RSNV) are included as tentative species (Brunt, 1995). Nicotiana velutina mosaic virus (NVMV), previously included as a tentative species, now has been removed from this genus. Recently, the re-classification of genus furovirus has been proposed into a family with three genera with SBWMV, BNYVV and PMTV as type species of each genus (Torrance and Mayo, 1997). SBWMV, OGSV, RSNV, SgCSV infect cereals, BNYVV, BSBV, PMTV infect root or tuber crops while BBNV, PCV infect leguminous crops. Eight viruses have plasmodiophorid 'fungus' vectors in genera, Polymyxa and Spongospora (Table 2).
Some of the properties of the furoviruses has been reviewed in recent past (Adams, 1988; Brunt, 1988; Cooper and Asher, 1988; Brunt and Richards, 1989; Campbell, 1996). This paper reviews briefly the information currently available on biology, transmission properties and mechanism involved in transmission process, molecular biology of furoviruses and also highlights exploitation of molecular pathological strategies for the protection of agricultural and horticultural crops against furoviruses.
Furoviruses
Taxonomy and Natural Occurrence: The group name, furovirus, for fungus-transmitted, rod-shaped plant viruses was first proposed by Shirako and Brakke (1984a) and was approved by the International Committee on Taxonomy of Viruses (ICTV) in 1987 (Brown, 1989). Furoviruses possess a divided genome consisting of two or more components of single-stranded ribonucleic acid (ssRNA) and are transmitted, through moist soil by motile zoospores of plasmodiophorid fungi. Previously, these viruses were considered as possible members of the monopartite tobamovirus group, based on particle morphology, serology, physico-chemical and biological properties (Fenner, 1976). However, virus particle instability, possession of fungal vector and their multipartite genome led to the current taxonomic placement (Shirako and Brakke, 1984a; Brunt, 1995). Eleven viruses have been classified as definite or possible members of the furovirus group (Brunt, 1991) (Table 1). They include four cereal-infecting viruses: soil-borne wheat mosaic virus (SBWMV), oat golden stripe virus (OGSV), rice stripe necrosis virus (RSNV) and sorghum chlorotic spot virus (SgCSV). Beet necrotic yellow vein virus (BNYVV), the causal agent of rhizomania disease of sugarbeet; beet soil-borne virus (BSBV) and potato mop-top virus (PMTV) infect root and tuber crops. Three infect legumes: broad bean necrosis virus (BBNV), peanut clump virus (PCV) and Indian peanut clump virus (IPCV). Hypochoeris mosaic virus (HyMV) and Nicotiana velutina mosaic virus (NVMV, Randles and Rohde, 1990) infect weeds. They have no known vector but possess other properties similar to those of furoviruses (Brunt, 1988) (Table 3). Sorghum chlorotic spot virus (SgCSV) is only serologically related to SBWMV but no other serological relationships have been reported.
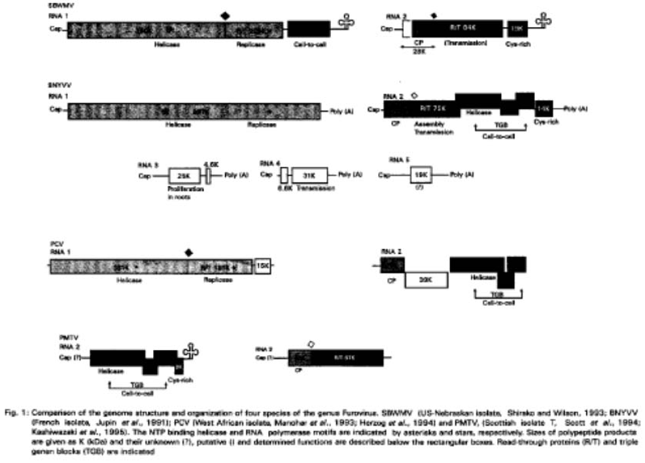
Classification and nomenclature of the furovirus group is still ambiguous. SBWMV (the type species) and BNYVV have a similar particle morphology, are transmitted by fungal vectors belonging to the Polymyxa genus and possess a divided RNA genome with the two largest RNA species carrying the essential genetic information. However, it has been shown that field isolates of BNYVV contain 4 to 5 distinct RNA species (Koenig et al., 1986; Kuszala et al., 1986; Tamada et al., 1989). RNA 3 of BNYVV promotes the systemic spread of virus in the root system and RNA 4 (Tamada and Abe, 1989) and the coat protein/read-through domain of RNA 2 (Tamada and Kusume, 1991) are associated with fungus transmission (Fig. 1; Table 4). The role of RNA 5 is unknown (Tamada et al., 1989; Jupin et al., 1991). In contrast, the SBWMV genome is bipartite (Shirako and Wilson, 1993) and differs from BNYVV in that (a) the 3'-end of SBWMV RNA probably forms a tRNA-like structure (Shirako and Wilson, 1993) whereas that of BNYVV is polyadenylated (Putz et al., 1983) and (b) SBWMV may have a cell-to-cell movement protein of the dianthovirus-type encoded at the 3'-end of RNA 1 (Shirako and Wilson, 1993), whereas BNYVV has triple gene block-type proteins encoded between the capsid protein gene and cysteine-rich protein gene in RNA 2 (Gilmer et al., 1992). PMTV has a tripartite genome and is not polyadenylated at the 3'-end (Reavy et al., 1993; Scott et al., 1994). The sequences of PMTV RNA 3 (Kashiwazaki et al., 1995) and RNA 2 (Scott et al., 1994) show that PMTV has a different genome organization within the genus Furovirus (Fig. 1). The arrangement of genes in PMTV RNA 3 resembles that of the 5' half of BNYVV RNA 2 and the arrangement of genes in PMTV RNA 2 resembles that of the 3' half of BNYVV RNA 2 (Kashiwazaki et al., 1995). PMTV coat protein is more similar to that of SBWMV than of BNYVV and the PMTV triple-gene-block proteins are more like those of barley stripe mosaic virus (BSMV) than of BNYVV (Scott et al., 1994). However, BSMV is a hordeivirus and has another kind of genome organization (Francki et al., 1991). Brunt (1995) has suggested that SBWMV, ORSV, PCV, PMTV and SgCSV be included as species of furovirus genus and BNYVV, BSBV, BBNV, HyMV, RSNV be included as tentative species of the genus. Since there is much heterogeneity among the member species, therefore, the taxonomy of the furoviruses should be revised (Shirako and Wilson, 1993; Herzog et al., 1994; Scott et al., 1994; Kashiwazaki et al., 1995). Recently, the re-classification of the genus furovirus has been proposed to be upgraded into a family with three genera while SBWMV, BNYVV and PMTV be the type species of the each genus (Torrance and Mayo, 1997). Recently, complete sequence of BBNV (Lu et al., 1998) and RNA 2 of BSBV (Koenig et al., 1997) suggested that both viruses are closely related to PMTV. These studies further support the idea of ''Furovirdae'' with three genera by including SBWMV, BNYVV and PMTV as type species.
Biological Properties of Furoviruses
Soil-Borne Wheat Mosaic Virus: Soil-borne wheat mosaic virus (SBWMV) is the type species of the genus Furovirus (Brunt, 1991, 1995). It was first recorded in Illinois and Indiana in 1919 and it has been found to occur in most winter wheat growing regions of the United States. These viruses has been reported from France, Italy, Japan, China, Egypt and Brazil (Table 1). In wheat, light green to yellow mosaic symptoms first appear in spring, occasionally in autumn and symptoms develop most frequently in plants growing in low lying areas of the fields. Such affected areas are often apparent from a distance, especially during extended cool spring weather but symptoms tend to be less conspicuous as plants mature and the average temperature increases (Koehler et al., 1952; Nykaza et al., 1979). SBWMV causes substantial yield losses each year (Brakke, 1971). The virus is sap transmissible but with difficulty and to only a few species of the Gramineae and Chenopodiaceae. In addition to wheat and barley, SBWMV infects Secale cereale, Bromus commutatus, B. tectorum and Chenopodium species.
Potato Mop-Top Virus: Potato mop-top virus (PMTV) was first detected by biological assay on seed potatoes growing on a farm at Auchterarder, Perthshire, Scotland, U.K. (B. D. Harrison, personal communication). Later an isolate (PMTV-T) was obtained from seed potatoes (Todd, 1965) and its soil-borne nature was determined in 1966 (Calvert and Harrison, 1966). PMTV was supposed to be transmitted by the potato powdery scab fungus, Spongospora subterranea (Wallr.) Lagerh. (Jones and Harrison, 1969) which was confirmed by acquisition and transmission of the virus in vitro by single cystosorus culture of the fungus (Arif et al., 1995). The virus causes a significant decrease in the yield and the quality of the tubers of susceptible potato cultivars and occurs in Northern and Central Europe, the Andean region of South America, Africa, Israel and Japan (Jones, 1988) and Canada (J. MacDonald, personal communication) (Table 1). Plants grown from PMTV-infected tubers produce yellow leaf blotching, shortening of internodes (Harrison and Jones, 1970; Cooper and Harrison, 1973; Harrison, 1974) and arched or distorted tubers but the type of symptoms induced depends greatly on the cultivar and environmental conditions. Some PMTV symptoms can be confused with those induced by other viruses e.g. aucuba on potato leaves by aucuba mosaic virus (Kassanis et al., 1972) and spraing in tubers, induced by tobacco rattle virus (TRV) (Harrison and Jones, 1970; Torrance, 1992) creating difficulties in diagnosis of PMTV. Both TRV and PMTV have rod-shaped particles and are soil-borne. However, TRV is a the type-member of the tobraviruses group and is transmitted by trichodorid nematodes in which it can persist for weeks to months. In contrast, PMTV is transmitted by S. subterranea (Arif et al., 1995) and persists in cystosori for more than twelve years (Jones and Harrison, 1969).
Previously, PMTV was classified as a possible member of the tobamovirus group (Kassanis et al., 1972) on the basis of particle morphology, coat protein size and weak serological relationship to tobacco mosaic virus (TMV). Subsequently, Randles et al. (1976) showed by immunoprecipitation tests that PMTV is serologically more closely related to SBWMV. In 1987, the ICTV approved the group name furovirus and PMTV was placed in the genus Furovirus on the basis of its tripartite genome and possession of a fungal vector (Brown, 1989; Brunt, 1995). Torrance and Mayo (1997) proposed re-classification of Furovirus and PMTV was included as type species of Pomovirus genus.
Peanut Clump and Indian Peanut Clump Viruses: The peanut clump disease, now known to be caused by peanut clump virus (PCV), was originally described from the Indian Subcontinent (Sundararaman, 1927). A similar disease was described in West Africa (Trochain, 1931) and was later found to be caused by a soil-borne virus (Thouvenel et al., 1976). PCV is believed to be transmitted by the plasmodiophorid fungus Polymyxa graminis (Thouvenel and Fauquet, 1981). A disease of peanut in India was originally considered to be caused by a distinct virus designated Indian peanut clump virus (IPCV) because it was apparently serologically unrelated to West African PCV (Reddy et al., 1979, 1983). Some authors have regarded IPCV and PCV as strains of one virus (Brunt and Richards, 1989) but the nucleotide sequence variation confirm that IPCV and PCV are distinct viruses (Wesley et al., 1994).
PCV has a wide experimental host range. Isolates from West Africa are transmissible by soil-bait tests to 39 species in eight families and those from India to between 13 and 20 species in Chenopodiaceae, Leguminosae and Solanaceae. It induces chlorotic and necrotic lesions in the inoculated leaves of Chenopodium quinua, Caravelin ensiforamis, Cyamopsis tetragonaloba, Phaseolus vulgaris and Vigna unguiculata and chlorosis or necrosis of systemically infected leaves of Capsicum annuum, Cassia occidentalis, Crotalaria juncea, Nicotiana benthamiana, N. clevelandii and N. edwardsonii (Thouvenel et al., 1976; Brunt and Richards, 1989). PCV is seed-borne in A. hypogaea but not seed-borne in great millet (S. arundinaceum) or N. benthamiana (Thouvenel et al., 1976). The natural occurrence of PCV in S. arundinaceum and other members of the Gramineae suggests that, though here considered to be a virus of a leguminous crop, it might well be recognized as yet another virus of cereals (Brunt and Richards, 1989).
Oat Golden Stripe Virus: The oat golden stripe virus (OGSV) was first recorded in infected plants displaying yellow stripe symptoms on the leaves in a crop of winter oats (Avena sativa) from Devon, South West England in 1967 (MacFarlane et al., 1968; Plumb and MacFarlane, 1977). It has been found also in West Wales (Catherall et al., 1977). Meanwhile a similar virus has also been detected in France since 1982, where the name soil-borne oat stripe virus is preferred (Brunt and Richards, 1989). The virus is occasionally found in plants naturally infected with oat mosaic virus (OMV), which has flexuous rod-shaped particles and is possibly a bymovirus. Single infection by OGSV results in chlorotic striping in the flag leaves of plants grown at temperatures of 15-20°C.
The virus is soil-borne and transmitted by the plasmodiophorid 'fungus', P. graminis which is also the vector of OMV and SBWMV (Plumb and MacFarlane, 1977). OGSV is mechanically transmissible from infected to healthy oat cultivar Blenda (Plumb and MacFarlane, 1977) and also induces chlorotic or necrotic local lesions in C. amaranticolor, N. debneyi and N. clevelandii (Plumb and MacFarlane, 1977; Brunt and Richards, 1989). Adams et al. (1988a) purified and determined the physico-chemical properties of the virus and also reported serological relationships to other species of the genus Furovirus. OGSV is serologically related to SBWMV but not to BNYVV and PMTV.
Sorghum Chlorotic Spot Virus: A virus which is morphologically and physico-chemically similar to SBWMV was isolated from Sorghum bicolor. The name sorghum chlorotic spot virus (SgCSV) was proposed for this virus because it produced dominant symptoms of elongated chlorotic spots and ringspots as well as yellowing on systemically infected leaves of sorghum and inbred Zea mays lines. SgCSV is mechanically transmissible to several monocotyledonous and dicotyledonous hosts and produces symptoms on the inoculated leaves of N. clevelandii, C. amaranticolor and C. quinoa.
The virus is bipartite with rods, 20 nm in diameter and of two modal lengths, 260 and 140 nm. Virions are composed of a single 20.5K capsid protein and two non-homologous, non-polyadenylated genomic RNAs of approximately 6.2 kb for RNA 1 and 3.5 kb for RNA 2. SgCSV and SBWMV capsid proteins are serologically related but no similarity exists between SgCSV RNA and SBWMV RNA, under high stringency conditions (Kendall et al., 1988).
Beet Necrotic Yellow Vein Virus: Beet necrotic yellow vein virus (BNYVV) is a fungus-transmitted soil-borne virus and is the causal agent of rhizomania disease of sugar beet (Beta vulgaris L. var. Saccharifera). The virus (BNYVV) and the disease (rhizomania) occur in most sugar beet growing areas of Europe, Asia and USA (Table 1). It is economically very important due to its severe effect on yield of the crop and quality of sugar (Putz et al., 1988).
The virus produces most conspicuous symptoms in sugarbeet roots, with stunting of infected plants and profuse development of lateral roots, giving them a bearded appearance, from which the name rhizomania is derived. In severely affected plants, the tap roots are stunted, lateral roots are killed and the vascular tissue is discoloured and necrotic. Leaf symptoms occur rarely, but leaf veins, sometimes, become chlorotic and necrotic (Brunt and Richards, 1989). BNYVV is transmitted in a persistent manner by the fungus Polymyxa betae Keskin (Abe and Tamada, 1986) which is a plasmodiophorid and obligate parasite of root tissue of a number of species (mostly Chenopodiaceae) including sugar beet.
Beet Soil-Borne Virus: Beet soil-borne virus (BSBV) was first recorded in England (Ivanovic et al., 1983) and it has furovirus-like particles and a fungal vector (P. betae). The virus is mechanically transmissible to C. album, C. amaranticolor, C. quinoa, Spinacia oleraceae and Beta vulgaris var. Saccharifera (Henry et al., 1986). BSBV and BNYVV are serologically different and no cross-hybridization have been observed between their RNAs or cDNAs (Lesemann et al., 1989).
Lesemann et al. (1989) and Hutchinson et al. (1990) reported a serological relationship between German BSBV isolates (Ahlum and Wierthe) and British and Swedish isolates, but no serological reaction has been found between any of these isolates and BNYVV. BSBV is a possible species of the genus Furovirus (Hutchinson et al., 1990; Kaufmann et al., 1992; Brunt, 1995).
Eight virus isolates with rigid, rod-shaped particle morphologies similar to BNYVV have been isolated from sugarbeet roots in the United States and referred as beet soil-borne mosaic virus (BSBMV, Liu and Duffus, 1988; Wisler et al., 1993a, b). Polyclonal antisera to the C-terminal 60 amino acids of to BNYVV CP weakly reacted with the 24K CP of BSBMV and reacted strongly with the 22K CP of BNYVV. No cross-reactivity was observed in reciprocal immunodiffusion tests between BNYVV and the BSBMV isolates, using antisera to CP of each virus. No products were observed for the BSBMV isolates when analysed in PCR using ten BNYVV primers pairs. All eight BSBMV isolates induced symptoms different from those of BNYVV in several hosts and two of them, tested, were transmitted by P. betae. It was concluded that these eight BSBMV isolates appear to be furoviruses distinct from BNYVV (Wisler et al., 1994) and may have an association with BSBV.
Broad bean Necrosis Virus: Very little information is available about broad bean necrosis virus (BBNV). The virus was first reported from Kyushu, Japan, from broad bean plants with veinal chlorosis and chlorotic spots on leaves. Plants infected in early spring were severely stunted and leaves of infected plants develop chlorotic vein spotting and older leaves had reddish-brown spots. Plants, infected, later were less seriously infected (Brunt and Richards, 1989). The virus is sap transmissible to 10 of 35 species in four of nine families (Inouye and Asatani, 1968; Inouye and Nakasone, 1980). At temperatures below 20°C, it induces systemic infection in Lathyrus odoratus, Pisum sativum and Vicia faba. C. amaranticolor and C. quinoa are useful diagnostic hosts for the virus (Brunt and Richards, 1989). BBNV is thought to be fungus-transmitted by Polymyxa spp. (Fujikawa, 1963) and is included in the genus Furovirus as a tentative species (Brunt and Shikata, 1986; Brunt, 1991, 1995).
Nicotiana velutina Mosaic Virus: Nicotiana velutina mosaic virus (NVMV) was first isolated from a Nicotiana velutina plant with bright yellow mosaic symptoms in the semi-arid zone of South Australia (Randles et al., 1976; Randles, 1978). It is mechanically transmissible to twenty species in the Solanaceae and Chenopodiaceae and also to Gomphrena globosa (Amaranthaceae); its host range therefore closely resembles that of PMTV. The rod-shaped NVMV particles fragment during purification and although particles up to 700 nm long have been reported (Randles et al., 1976) the modal length of partially purified particles is 125 to 150 nm. It has no known vector but has a high rate of seed transmission in several Nicotiana sp. On the basis of particle morphology (a helical capsid of 2.9 nm pitch with a diameter of 18 nm) and coat protein 21.4 K (Randles et al., 1976), NVMV can be placed in any of the tobamovirus, hordeivirus, tobravirus, (Matthews, 1982) or furovirus genera (Brunt and Shikata, 1986; Brunt and Richards, 1989; Brown, 1989; Brunt, 1995). No serological relationship has been found between NVMV and PMTV, BSMV, BNYVV, SBWMV, TMV or TRV (Randles et al., 1976). Randles and Rohde (1990) reported that NVMV has affinities with members of the hordeivirus, furovirus, potexvirus and carlavirus groups but not with the tobravirus group. The divided RNA genome of NVMV and the sizes of both of its two RNAs, suggest that NVMV is most closely related to furoviruses (Brunt, 1991).
Hypochoeris Mosaic Virus: Hypochoeris mosaic virus (HyMV) was found in Hypochoeris radicata, a weed host near Vancouver, British Columbia, Canada (Brunt and Stace-Smith, 1978a). A similar virus was found in Leontodon autumnale, a common weed in potato growing areas of Eastern Canada and also in Hypochoeris radicata in Australia (Greber and Finlay, 1981). This virus infects a large number of host plants but there is no evidence yet available about its natural vector.
The virus has rod-shaped particles with modal lengths in sap of 120 to 140 nm and 220 to 240 nm, very similar to those of SBWMV, PMTV, BNYVV, BBNV, NVMV, PCV, BSMV and the defective strains of TMV. It is serologically and pathogenically distinct from all of these viruses (Brunt and Stace-Smith, 1978a, b). It was classified as a possible member of the tobamovirus group (Brunt and Stace-Smith, 1978a) but now it is included as a tentative species of the genus Furovirus (Brunt, 1995).
Rice Stripe Necrosis Virus: Rice stripe necrosis virus (RSNV) is transmitted by P. graminis. The virus produces chlorotic or necrotic stripes on the leaves of infected plants. It is not transmitted mechanically to rice, but it does induce lesions in mechanically inoculated leaves of C. amaranticolor and N. benthamiana (Brunt and Richards, 1989; Fauquet et al., 1988).
General Characteristics of Furoviruses
Particle Morphology, Structure and Composition: Furovirus particles consist of hollow rigid nucleoprotein rods about 20 nm in diameter, with two or more modal lengths (270 and 150 nm). The predominant bimodal particles suggest that they have bipartite genomes, but this has yet to be demonstrated for some viruses (Brunt, 1995). Both particles are needed for infection of PCV (Reddy et al., 1985), PMTV (Randles et al., 1976), SBWMV (Shirako and Brakke, 1984a) and BNYVV (Putz et al., 1983). In BNYVV and RSNV the long and intermediate particle lengths are 390 and 270 nm, respectively (Tamada, 1975; Putz, 1977; Fauquet et al., 1988). Particles up to 700 nm in length have been reported for NVMV but the range is extremely wide with a modal length of about 125-150 nm (Randles et al., 1976; Randles and Rohde, 1990) (Table 3). PMTV has fragile rod-shaped particles, 18-20 nm wide and with two predominant lengths, 100-150 nm and 250-300 nm (Harrison and Jones, 1970; Kashiwazaki et al., 1995). Furovirus particles contain many copies of a single polypeptide, with a molecular weight between 19 and 24K, assembled as right handed single-start helices. PMTV particles are helically constructed with an estimated pitch of 2.4-2.5 nm (Harrison and Jones, 1970; Kassanis et al., 1972). Kallender et al. (1990) reported that PMTV contains three ssRNAs of 6.5 kb, 3.2 kb and 2.5 kb. Recently, Scott et al. (1994) reported the presence of three species of distinct RNAs with sizes of 6.4 kb, 3.0 kb and 2.5 kb. The protein in PMTV particles consists of a single polypeptide species of molecular mass 19.7 K (Torrance et al., 1993; Kashiwazaki et al., 1995; Arif and Reavy, 2000). Optical diffraction of BNYVV particles has shown that the protein helix is right handed with pitch of 2.6 nm and 12.5 protein subunits per turn (Steven et al., 1981). The RNA comprises about 5% of the particle weight. No data are available on the presence of divalent cations, ubiquitinated coat protein subunits or polyamine content (Shirako and Wilson, 1994).
Serological Properties: Most species of the genus Furovirus are good immunogens. Tests, with polyclonal antisera, showed that PMTV, SBWMV, OGSV and BBNV are serologically distantly related (Kassanis et al., 1972; Randles et al., 1976; Nakasone and Inouye, 1978). Also SBWMV and PMTV are reported to be distantly related to tobacco mosaic virus (TMV) (Kassanis et al.,1972; Powell, 1976; Fenner, 1976). BBNV is unrelated to TMV (Nakasone and Inouye, 1978) but is related to two other tobamoviruses, cucumber green mottle virus (CGMV) and odontoglossum ringspot virus (ORSV) (Shirako and Wilson, 1994).
Molecular Biology of Furoviruses: In the following section the molecular biology of four well characterized species in the genus have been briefly discussed. Molecular studies of the SBWMV genome indicated that the lengths of RNA 2 is variable. The length of RNA 1 has been estimated to be 7098 nt (Shirako and Brakke, 1984a) and RNA 2 is 3592 nt (Shirako and Brakke, 1984a, b; Shirako and Wilson, 1993) but the length of RNA 2 depends on the growing conditions of the infected wheat plants. RNA 2, isolated from naturally infected wheat [referred to as wild type (WT)], is about 3500 nt but when the same RNA species is isolated from manually-inoculated wheat plants and kept for 2 years at 15-17°C, the size is 2800 nt (700 nt shorter than WT RNA 2 and referred as Lab. 2). The size of RNA 2 also varies from plant to plant or even different parts of the same plant when samples are taken at different times during the growing season (Shirako and Brakke, 1984a, b; Hsu and Brakke, 1985a, b; Shirako and Ehara, 1986). The deleted RNA 2 molecules are infectious when co-inoculated with RNA 1 and induce more severe symptoms than full length RNA 2 molecules (Shirako and Ehara, 1986). Chen et al. (1994) detected 1058 and 759 nt deletions in RNA 2 of SBWMV-isolates Lab-1 and Okl-7, respectively, within the region encoding the coat protein read-through domain.
Both RNAs (RNA1 and RNA2) of SBWMV have a 5' cap structure (m7GpppG) but are not polyadenylated at the 3'-end. Instead, the 3' termini are either -CCOH or -CCAOH and the last 50-100 nucleotides have similarity with the corresponding region in sunn-hemp mosaic virus (SHMV, a tobamovirus) and turnip yellow mosaic virus (TYMV a tymovirus). The 3' region may form a functional tRNA-like structure using a series of pseudoknots as in TMV RNA (Shirako and Wilson, 1994). Shirako and Wilson (1994) further reported that SBWMV RNA 1 has 101-nt of untranslated sequence before the first AUG codon of an open reading frame (ORF) which encodes a 150K polypeptide. This ORF ends with an opal termination codon which can be suppressed (at least in vitro) to extend the 150K polypeptide to give a 209K protein which has similarity with the equivalent 126K protein of TMV and tobacco rattle tobravirus (TRV) RNA 1. Following the 209K ORF there is an ORF for a polypeptide of 34K which shows weakly pregnant with the TMV 30K and TRV RNA 1 29K proteins associated with cell-to-cell movement. In contrast, the cell-to-cell movement proteins of BNYVV consist of a triple gene block, similar to those of potexviruses, carlaviruses and hordeiviruses and are encoded on RNA 2. The genome organization of SBWMV RNA1 most closely resembles TMV RNA (but for the absence of the 3'-ORF for coat protein) and TRV RNA 1 (except for the absence of the 3'-ORF for a cysteine-rich 16K protein), shares little similarity with BNYVV RNA 1 (Shirako and Wilson, 1994).
SBWMV RNA 2 has a 333 nt untranslated leader sequence before the first large ORF which encodes the 19K coat protein (CP). Another AUG codon in the same reading frame as the true CP start codon occurs in the RNA 2 leader at position 100-102. Translation of transcripts from deleted clones shows that this AUG is not functional (Hsu and Brakke, 1985a, b; Shirako and Wilson, 1993, 1994). Following the termination codon for the protein is an in-frame ORF for a potential 83K protein resulting from read-through of this termination codon. Shirako and Ehara (1986) compared SBWMV to a Japanese isolate of BNYVV and showed that the 83K read-through protein has sequence similarity with the 75K polypeptide of BNYVV.
The nucleotide sequences of PMTV RNA 2 (2962 nt) and RNA 3 (2315 nt) have been determined (Scott et al., 1994; Kashiwazaki et al., 1995). RNA 2 contains four overlapping ORF's encoding (from 5'-end) polypeptides of molecular weight 51K, 13K, 21K and 8K. The first three ORFs constitute a triple-gene-block which is homologous to that of other rod-shaped and elongated viruses; particularly barley stripe mosaic hordeivirus. The 51K protein contains a putative NTP-binding motif and the 13K and 21K protein each contain two hydrophobic regions, separated by a hydrophilic region (Reavy et al., 1993; Scott et al., 1994). The 8K protein is rich in cysteine. RNA3 contains the virus coat protein gene (20K) and associated read-through protein (47K) which may play a role in transmission by the fungal vector (Reavy et al., 1993; Kashiwazaki et al., 1995).
PCV contains two genomic RNAs of 5900 nt and 4300 nt, respectively (Reddy et al., 1985). Neither RNA 1 nor RNA 2 is polyadenylated at its 3'-end. The structure of the 5'-end is unknown, but it is unlikely to contain a genome linked protein (VPg) since proteinase K has no effect on the infectivity of PCV RNA. However, the presence of VPg is not an absolute requirement for infectivity of the RNA of some plant viruses (Mayo et al., 1982). As with SBWMV RNA 1, PCV RNA 1 encodes a major protein of 143K and a minor (read-through) product of 195K in vitro. RNA 2 directs the synthesis of the 24.5K coat protein but no read-through product is made in vitro. Products of 50K and 20K also appeared from minor PCV-related RNAs and it is not known whether these species are sub-genomic RNA derived from RNA 1 or RNA 2 are satellite RNAs (Reddy et al., 1985). Manohar et al. (1993) determined that the complete 4503-nucleotide sequence of PCV RNA 2 contains six ORFs. The ORF1 located nearest to the 5' end of the RNA contains the virus coat protein gene (23K) and is separated from the 5-'extremity by a non coding region of 389 nucleotides. The AUG codon of ORF 2 overlaps with the UGA termination codon of ORF 1 so that ORF 2 starts one nucleotide upstream from the first residue of the termination codon of the coat protein cistron and encodes a polypeptide of 39K. Three additional ORFs with the potential to encode polypeptides of approximately 51K, (ORF 3), 14K (ORF 4) and 17K (ORF 5) are found in the 3' end of RNA 2 (Herzog et al., 1994, 1995). ORFs 3, 4, 5 resemble the triple gene block of potexviruses, carlaviruses, barley stripe mosaic hordeivirus and beet necrotic yellow vein furovirus. The 276 nt 3' noncoding region of RNA 2 displays sequence similarity with the 3' non-coding region of RNA 1.
The BNYVV genome contains 4-5 RNA molecules. Sequence analysis indicated that sizes of RNAs are 6746 nt (RNA 1), 4612 nt (RNA 2), 1774 nt (RNA 3) and 1467 nt (RNA 4) (Bouzoubaa et al., 1985, 1986, 1987). Recently RNA 5 (1400 nt) and RNA 6 (1000 nt) have been reported in some Japanese isolates of BNYVV. RNA 5 is distinct from other genomic species (RNA 1, 2, 3, 4) in BNYVV and RNA 6 is a deletion mutant of RNA 5 (Kiguchi et al., 1989). The status of these RNAs is uncertain as they are present in some but not all Japanese isolates of BNYVV but have not yet been found in isolates from other parts of the world (Tamada et al., 1989; Richards and Tamada, 1992). RNA 1 and RNA 2 are needed for virus infection, whereas the smaller species, RNA 3, RNA 4 and RNA 5, behave like satellite RNA molecules in mechanically inoculated leaves of host plants (Bouzoubaa et al., 1985; Koenig et al., 1986; Tamada et al., 1989; Jupin et al., 1990a; Bouzoubaa et al., 1991). RNA 3 and RNA 4 may undergo deletion or disappear when the isolates are passed serially by mechanical inoculation of C. amaranticolor (Bouzoubaa et al., 1986). It has also been found out that RNA 3 greatly affects symptom expression in host plants (Kuszala et al., 1986; Tamada and Abe, 1989; Tamada et al., 1990; Jupin et al., 1990b, 1992) and RNA 3 is responsible for rootlet proliferation and facilitates viral spread, in root tissues. RNA 3 may also confer some properties that are necessary for virus survival in the field (Tamada and Abe, 1989). Lemaire et al. (1988) reported that full-length RNA 3 and RNA 4 seem to play a role in transmission by the fungal vector. However, it was unclear whether RNA 3 or RNA 4, or both, are needed for fungal transmission. Tamada and Abe (1989) described the results of transmission tests using virus isolates containing different combinations of small RNA species and reported that RNA 4 plays an important role in fungal transmission. Evidence was presented that the 75K read-through product encoded by the second ORF of RNA 2 is essential for transmission of BNYVV by P. betae (Tamada and Kusume, 1991) and is also important in virus assembly (Schmitt et al., 1992). A summary of gene functions, among well characterized furoviruses, are given in Table 4 and illustrated in Fig. 1. Recently, sequence of RNA 2 of Ahlum isolate of BSBV have been reported (Koenig et al., 1997). BBSV RNA 2 was similar in genetic organization to PMTV RNA 3, although it was 1100nt longer. Its 3'-end, unlike that of PMTV RNA 3, had the potential to fold into a tRNA-like structure. It contains one large ORF for RT protein with a molecular mass of 104K which was interrupted internally by an amber stop codon terminating the coding region for CP of 19K. The RT domain was much longer than that of PMTV. Recently, Lu et al. (1998) reported the sequence of BBNV-O, a tripartite genome (RNA 1 6.0kb; RNA 2 2.8kb; RNA 3 2.4kb). RNA 1 contains methyltransferase, helicase and RNA-dependent RNA polymerase (RdRp) motifs. The CP (20K) cistron was located at the 5'-terminal end of RNA 2 and partially produce a large RT protein. RNA 3 possesses typical motifs of triple gene block proteins. The genome organization and phylogenic analysis, using RdRp and CP amino acid sequence, suggested that BBNV is closely related to PMTV and distantly related to other furoviruses.
Fungal Transmission: The viruses with fungal vectors can be divided on the basis of particle morphology and genome organization into three different types: 1. isometric viruses (carmoviruses, necroviruses and some uncharacterized agents) which are transmitted by chytridiomycete fungi (Olpidium species); 2. straight, tubular rods (furoviruses); and 3. flexuous, filamentous particles (bymoviruses). The furo- and bymoviruses are transmitted by plasmodiophorid fungi in the species, Polymyxa and Spongospora (Table 5).
Plasmodiophorid Fungi: Taxonomy and Life Cycle: Fungi, known to transmit furoviruses, are zoosporic, obligate parasites of plant roots and belong to the class plasmodiophoromycetes, subdivision Mastigomycotina and division Amastigomycota (Adams, 1991a; Brunt, 1989). P. graminis and P. betae, generally not thought to be pathogenic to their hosts, are morphologically indistinguishable, but have different host ranges (Barr, 1979). S. subterranea is the causal agent of powdery scab of potato tubers (Hims and Preece, 1975; Teakle, 1983; Harrison et al., 1997) and crook root disease of watercress (Tomlinson, 1958; Clay and Walsh, 1990). Braselton (1988) and Barr (1988) have discussed the classification and life cycles of plasmodiophoromycetes. The life cycle of each species possess biflagellate zoospores, multinucleate plasmodia and thick-walled clusters of resting spores (cystosori). Cystosori are formed in plant roots and remain in the soil when the roots decay. These resting spores remain in the soil for several years and can even survive passage through the alimentary canals of herbivores. The primary zoospores, produced by germination of cystosori, swim to host root hairs (or epidermal cells on the surface of tubers) where they encyst and inject the protoplast contents into the host cell. The fungus produces a multinucleate plasmodium which develops into a mature zoosporangium. This sac-like structure liberates secondary zoospores, which are usually indistinguishable morphologically from primary ones, into soil water and these zoospores reinfect the plant producing more zoosporangia or cystosori (Braselton, 1988; Barr, 1988; Adams, 1991b). The optimum temperature for the growth of P. graminis and S. subterranea is 18°C and 15°C (Brakke, 1971; Jones and Harrison, 1972). Certainly SBWMV, BNYVV and PMTV become problems when soil temperatures are less than 18°C and 15°C, respectively and symptoms, in infected plants, become more conspicuous when temperatures do not exceed 20°C (Brakke, 1971; Jones and Harrison, 1972; Harrison, 1974). By contrast, PCV spreads efficiently in soils at 28°C or warmer climate in West Africa and India (Thouvenel et al., 1976; Mayo and Reddy, 1985).
Mixing suspensions of P. graminis zoospores and SBWMV virions do not produce a viruliferous vector (Rao and Brakke, 1969) implying that the virus persists inside the zoospores. Successful transmission of virus by zoospores, in the presence of homologous antiserum, has been reported for BaMMV (Adams and Swaby, 1988), BNYVV (Abe and Tamada, 1986) and SBWMV (Rao and Brakke, 1969). Virus particles are presumably carried internally, hence transmission cannot be affected by treating motile zoospores with antiserum or by washing cystosori with acid or alkali (Adams, 1991b).
Abe and Tamada (1986) detected BNYVV in immature zoospores within zoosporangia of P. betae, in section of infected sugarbeet roots by electron microscopy. However, attempts to locate SBWMV, BNYVV particles, in different life stages of the Polymyxa, have been unsuccessful (Langenberg and Giunchedi, 1982) and BaMMV (Langenberg and Van Der Wal, 1986). Immunogold labelling has been used to locate BaMMV within zoosporangia of P. graminis and in zoospores. However, only 1-2% of zoospores carried virus (Chen et al., 1994). The amount of virus in the fungus probably depends on the amount of virus in the host cell at the time of infection by the fungus as in some cases up to 50% of P. betae plasmodia were found to contain virus (Rysanek et al., 1992).
Types of Virus Transmission by Fungal Vectors: In the study of fungal vectors of plant viruses, two types of virus-vector relationship have been established (Campbell, 1979, 1993, 1996, Teakle, 1983; Adams, 1991b), one in which the viruses were acquired in vitro and survived externally on the resting spores and another relationship in which viruses were acquired in vivo and survived internally. These relationships were based on the facts whether the virus was acquired in vivo or in vitro and whether the virus survived externally or internally to the resting spores, in the absence of a living host plant (Campbell, 1993). The terminology of non-persistent and persistent was adopted from the work on aphid transmission (Watson and Roberts, 1939; Watson, 1960). Watson (1960) defined persistence as "the time for which a vector can go on infecting healthy plants after it leaves the infected ones" or more briefly as "persistence in feeding vector".
| Table 2: | Natural fungal vectors of furoviruses |
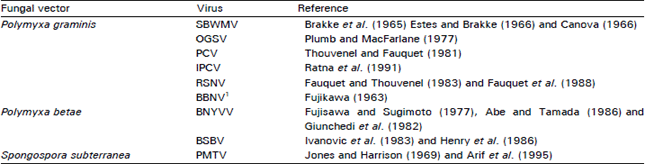 | |
| 1Polymyxa species has been reported as vector of the virus but further confirmation has not been made | |
| Table 3: | Particle size and genome composition of furoviruses |
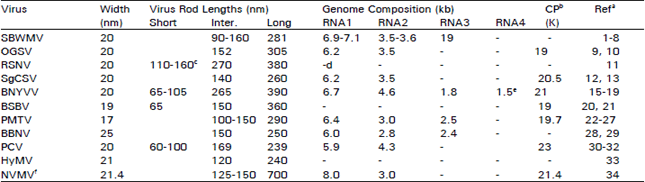 | |
a1Brakke (1971), 2Gumpf (1971), 3Tsuchizaki et al. (1975), 4Brakke (1977), 5Shirako and Brakke (1984a), 6Shirako and Ehara (1986), 7,8Shirako and Wilson (1993, 1994), 9Adams et al. (1988a), 10Shirako and Wilson (1994), 11Fauquet et al. (1988), 12 Kendall et al. (1988), 13Shirako et al. (1993), 14Tamada (1975), 15Putz (1977), 16,17,18Bouzoubaa et al. (1985, 1986, 1987), 19Richards and Tamada (1992), 20Kaufmann et al. (1992), 21Koenig et al. (1997), 22Roberts and Harrison (1979), 23Kallender et al. (1990), 24,25Scott et al. (1990, 1994), 26Kashiwazaki et al. (1995), 27Pereira et al. (1994), 28Inouye and Nakasone (1980), 29Lu et al. (1998), 30Thouvenel and Fauquet (1981), 31Reddy et al. (1985), 32Manohar et al. (1993), 33Brunt and Stace-Smith (1978a), 34Randles and Rohde (1990) | |
| bCapsid protein |
| cMay be fragments of the long or intermediate particles |
| dDenote parameter unknown |
| e1.4 kb RNA 5 and 1.0 kb RNA 6 have been reported in some Japanese isolates of BNYVV (Richards and Tamada, 1992) |
| fPreviously included as a tentative species of the genus, also has a bipartite genome (8 kb and 3 kb); the sizes of its two RNAs, however, differ from those of furoviruses and is is now probably best excluded from the genus (Brunt, 1995) |
| Table 4: | Functions of gene products of furovirus genomes1 |
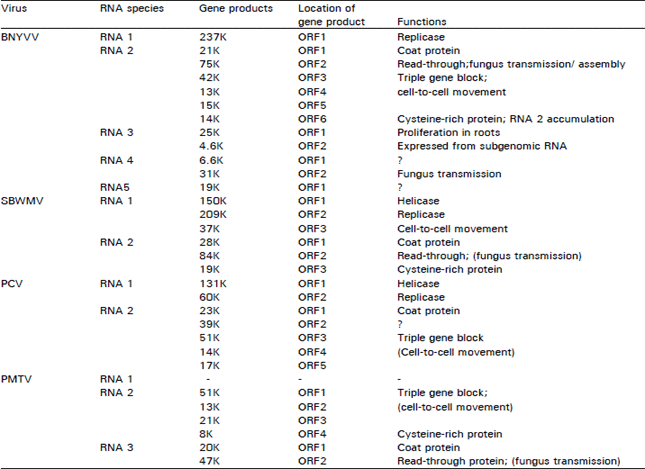 | |
1Sizes of polypeptide gene products are given in K (Kilodaltons); locations are shown on Open Reading Frames (ORFs) and their unknown (?); putative() or determined functions are given | |
| Table 5: | Transmission of plant viruses by soil-borne fungi |
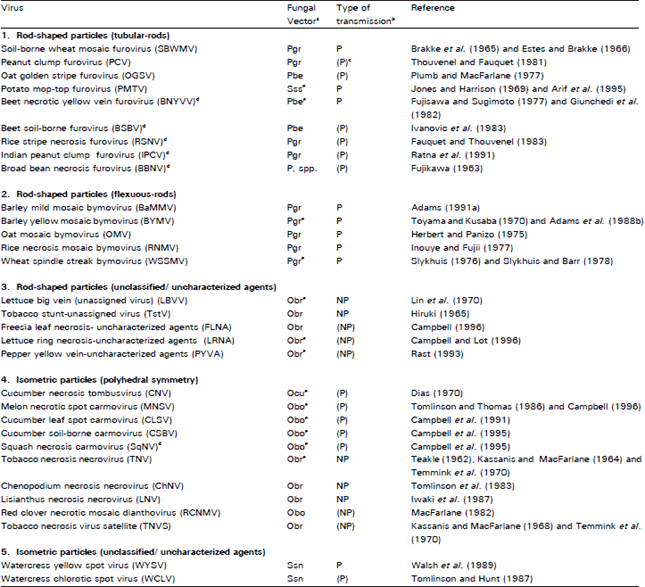 | |
aAbbreviations used for vectors: Pbe = Polymyxa betae Keskin, Pgr = Polymyxa graminis Ledingham; Sss = Spongospora subterranea (Wallr.) Lagerh f. sp. subterranea; Ssn = Spongospora subterranea (Wallr.) Lagerhf. sp. nasturtii; Obr = Olpidium brassicae (Wor.) Dang; Obo=Olpidium bornovanus = (O. radicale) Schwartz and Cook; Ocu = Olpidium cucurbitacearum Barr and Dias.; P. spp.= Polymyxa (species unknown). b P = Persistent: virus particles are carried inside the fungal zoospores but are not thought to be multiply within the vector; NP = Non-persistent: virus particles are carried on the surface of the zoospore. c Information in parentheses needs to be confirmed. d Tentative species of the genus (Murphy et al., 1995). e Virus transmission by fungal vector has been demonstrated through monofungal (unifungal/ single cystosorus) or equivalent culture. | |
Teakle (1983) reviewed fungal transmission of viruses and offered these definitions: ''persistent viruses are internally in zoospores and survive, often for many years, in dormant resting sporangia'', whereas the ''non-persistent viruses are borne on the surface of the zoospores and do not persist in the resting sporangia''. This terminology was used by Adams (1991b); where he r e viewed the mechanisms, involved in virus transmission in relation to the non-persistent and persistent manners. He used non-persistent transmission for tobacco necrosis virus (TNV) and later tobacco necrosis satellite virus (TNSV) and cucumber necrosis carmovirus (CNV) with relation to their fungal vector Olpidium spp. in which viruses could be acquired by zoospores in vitro and appear to be carried externally on the zoospores and resting spores. Persistent transmission was used for lettuce big vein virus (LBVV) and tobacco stunt virus, (TSV), because the virus was only acquired in vivo and was assumed to be carried internally. He differentiated persistent and non-persistent types of transmission, on the basis of the characteristics proposed by Campbell (1993) reviewed the use of Teakle (1983) which can be summarised as:
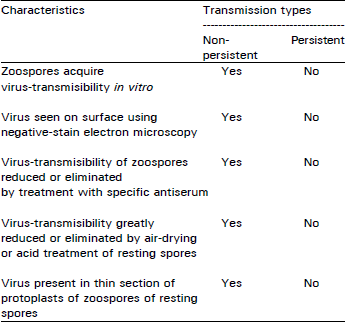
persistent and non-persistent terminology and suggested that the existing situation is ambiguous and misleading. He pointed out that virus acquired in vitro, can be taken into the zoospore protoplasm and thus the definitions are not absolute. He felt that a new terminology reflecting the unique aspects of fungus vector/ virus interactions should be devised instead of the use of terms derived from invertebrate vector. He suggested that the International Working Group on Plant Viruses with Fungal Vectors (IWGPVFV) should revise the terminology of fungal transmitted plant viruses.
Factors Controlling the Specificity of Virus Transmission: Furoviruses are acquired by fungi only in vivo, from infected root cells, by an unknown mechanism. It is known that once contact is made with a plant root, the zoospores become encysted and the zoospore contents (including the virions) enter the epidermal cell (Shirako and Wilson, 1994). The virus is probably released into the host cytoplasm at this time and starts its multiplication. The fungal plasmodium either becomes a sporangial plasmodium (or zoosporangium) from which viruliferous zoospores are released that infect additional root hairs and epidermal cells or becomes a cystogenous plasmodium that produces cystosori (Adams, 1991b). Arif (1995) proposed a mechanism by which virus-free zoospores of S. subterranea become viruliferous during acquisition and transmission process, through a series of events taken place in vivo. Virus-free cystosori germinate to produce virus-free zoospores which swim to susceptible host roots, encyst and penetrate into the host cytoplasm. The fungal mass then multiplies in the cytoplasm in the presence of virus particles and produces multinucleated fungal masses (sporangial plasmodia or zoosporangia) which yield secondary zoospores. It seems that virus particles, from the cytoplasm, are absorbed by the fungal plasmodium at this stage by phagocytosis or pinocytosis and during nuclear division, some of the secondary zoospores became viruliferous. The viruliferous secondary zoospores can then infect more roots and tubers and produce viruliferous plasmodia (cystogenous plasmodia) and subsequently produce clusters of viruliferous cystosori. The virus can persist in the cystosori for several years in moist soils. When these cystosori germinate they go on to infect subsequently virus-free plants (Fig. 2). This area of research has been almost completely neglected and needs to be exploited in order to design effective control strategies for both the virus and the vector.
Carmoviruses, dianthoviruses, necroviruses and associated satellites are transmitted by Chytridiomycetes fungi. Two species of Olpidium, O. brassicae (Wor.) Dang. and O. radicale Schwartz and Cook (= O. bornovanus) have been shown to transmit plant viruses (Table 4). Some of these viruses have isometric particles ranging in diameter from 17 nm (tobacco necrosis satellite, TNSV) to 31 nm (cucumber necrosis virus, CNV). Lettuce big vein virus (LBVV) and tobacco stunt virus (TSV) are serologically-related viruses that have rod-shaped particles 22 nm x 200-375 nm (Adams, 1991b). Most of the viruses, transmitted by Plasmodiophoromycetes, belong either to the furoviruses or bymoviruses. Furoviruses usually have particles of two or more lengths about 20 nm in diameter and up to 400 nm long (Table 3). Bymoviruses have flexuous, filamentous rods and have particles typically of two lengths (around 300 nm and 600 nm although those of WSSMV appear to be longer) and diameter of about 13 nm (Adams et al., 1989). The genomes of viruses transmitted by plasmodiophoromycetes are usually bipartite but BNYVV has four components and PMTV contains three distinct RNA components (see Fig. 1; Table 2). Bymoviruses have a bipartite, single-stranded, 3'-polyadenylated RNA genome (Kashiwazaki et al., 1990, 1991; Koenig and Huth, 1988; Davidson et al., 1991).
Role of Read-through Protein Gene in Transmission Process: In furoviruses read-through proteins are a common feature and are found in BNYVV (Bouzoubaa et al., 1986), SBWMV (Shirako and Wilson, 1993), PMTV (Kashiwazaki et al., 1995) and BSBV (Koenig et al., 1997). The BNYVV RT protein is involved in particle assembly (Schmitt et al., 1992) and transmission of the virus by its fungal vector, P. betae (Tamada and Kusume, 1991). Repeated manual passage of BNYVV results in spontaneous deletions of RT domainin, the part of RNA 2 which encodes the C-terminal region of the RT protein results in a loss of the ability of the virus to be transmitted by P. betae (Tamada and Kusume, 1991; Tamada et al., 1996). Spontaneous deletions have also been observed in the RT protein coding region of SBWMV RNA 2 (Shirako and Brakke, 1984b; Chen et al., 1995a, b) and PMTV RNA 3 (Reavy et al., 1998; Arif et al., 1999a) and deletions in SBWMV RNA 2 have been associated with an increase in symptom severity (Chen et al., 1994). Comparison of the RT protein of BSBV and PMTV (isolate T) indicated the presence of two domains (A and B) of conserved sequence near the N termini of the protein and a third domain (C) at the C-terminal end of the PMTV-T protein or 49 amino acids, from the C terminus of the BSBV protein (Koenig et al., 1997). The region between domains B and C of the PMTV-T RT protein was much shorter than that of the BSBV protein and was lacking in two regions of sequence conserved between BSBV and SBWMV. Recently it has been demonstrated that a field isolate of PMTV (PMTV-S; Arif et al., 1994) was efficiently acquired and transmitted by a monofungal culture of S. subterranea while PMTV-T could not be acquired and transmitted by the same fungal culture (Arif et al., 1995; Arif et al., 1999b). PMTV-T has been maintained by manual inoculation to N. debneyi or N. benthamiana for more than 30 years. Sequence analysis of RT protein coding region of PMTV-S showed the presence of an additional 543 nt in the 3' half of the coding region relative to that of PMTV-T.
 |
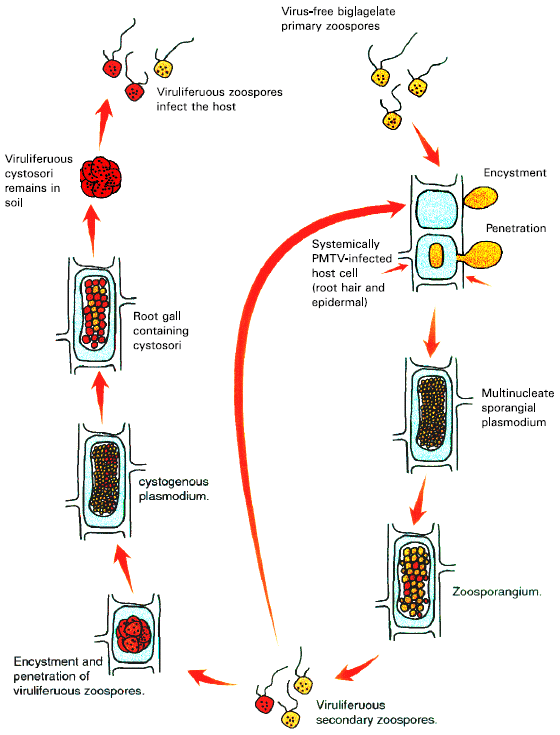 | |
| Fig. 2: | Illustration of life cycle of Spongospora subterranea with different hypothetical events in PMTV transmission |
These additional nucleotides preserved the reading frame of the RT protein and inserted 181 amino acids into the RT protein (Arif et al.,1995; Reavy et al., 1997). The association of lack of transmissibility of PMTV-T with an apparent deletion of sequence in RNA 3, relative to the RNA 3 of the transmissible isolate PMTV-S, suggests that the RT domain encoded by PMTV-S RNA 3 contains determinants that play an important role in acquisition and transmission of PMTV by S. subterranea.
Transmission studies, with bymoviruses (BaMMV and BaYMV) and P. graminis, indicated that an isolate of BaMMV from Streatly, Bedfordshire, UK, had lost the ability to be transmitted by P. graminis after repeated mechanical inoculation (Adams et al., 1988b). The complete genome of field isolates of BaYMV (Kashiwazaki et al., 1990; 1991; Davidson et al., 1991; Peerenboom et al., 1992) and BaMMV (Kashiwazaki et al., 1992; Foulds et al., 1993; Schlichter et al., 1993; Timpe and Kuhne, 1994) have been sequenced. Jacobi et al. (1995) reported the sequence of a BaMMV isolate which was not transmitted by P. graminis. A deletion of 1092 nt, at the 3'-terminal part of the RNA 2, was reported and it was suggested that the C-terminal protein plays an essential role in fungus transmission of BaMMV. Similar studies, with a French isolate of BaMMV, indicate that a 1000 nt deletion at the 3'-terminal part of RNA 2 controls transmission by P. graminis (Dessens et al., 1995). Sequence similarity between the bymovirus C-terminal protein (P2) and the capsid read-through protein, on furoviruses, suggests an evolutionary relation between bymo- and furoviruses (Dessens et al., 1995). McLean et al. (1994) assessed the role of cucumber necrosis tombusvirus (CNV) coat protein in the specificity of transmission by O. bornovanus. He constructed full-length infectious clones containing reciprocal exchanges between the coat protein gene of CNV and the cherry strain of tomato bushy stunt tombusvirus (TBSV-Ch). Virions containing TBSV-Ch genome with the CNV coat protein gene were efficiently transmitted, but those containing the modified CNV genome with TBSV-Ch coat protein gene were not. It is not known if the specificity of fungus/ virus reside in virus acquisition by the vector or in its transmission to the host (or both). It is expected that reverse genetics of infectious cDNA clones of these viruses and comparative transmission by their fungal vector may be able to elucidate determinant of transmission and answer these questions.
Virus Ecology and Control: Furoviruses naturally occur almost all over the world. Although some viruses have a restricted geographical distribution, others such as SBWMV, BNYVV, PMTV and HyMV are widely distributed. Because furoviruses are retained for many years in soil-borne resting spores of their fungal vectors, it is difficult to control the disease once a field becomes infested with viruliferous fungi (Cooper and Asher, 1988).
Conventional control measures including rotation of susceptible with fungus- or virus- resistant crops can decrease but not eliminate the amount of field inoculum. Delayed crop sowing in the autumn decreases the number of SBWMV-infected winter wheat plants, presumably because dry and warm climatic conditions prevent infection of roots by motile zoospores (Shirako and Wilson, 1994). The use of PCV-free groundnut seed, or high grade seed potatoes and rogueing of infected potato plants can decrease the incidence of PCV and PMTV (Germani et al., 1975; Cooper et al., 1976; Jones, 1988). Removal of weed hosts in which the fungus or virus overwinter is also beneficial. A single dominant gene from hard red winter wheat has been successfully used to delay SBWMV multiplication and accumulation until late in the season when symptoms are less pronounced even though the wheat plants are not completely resistant or tolerant to infection by either the fungus or the virus (Brakke and Langenberg, 1988). The incidence of PMTV and SBWMV can be reduced in small test plots by chemical sterilants and fungicides (methylbromide, formaldehyde, captan and benomyl) but these are expensive and ecologically undesirable for large-scale use (Solomon and Wastie, 1988; Jones, 1988; Usugi, 1988). Similarly, reduction of soil pH to 5.0 with sulphur, or addition of mercuric or zinc compounds, can kill zoospores but does not affect resting spores of the fungal vector and are also dangerous to the environment (Cooper et al., 1976).
Molecular Pathology: Despite advances in the production of virus-free seed stocks and the control of virus vectors by agrochemicals, virus infection still cause serious economic losses in agricultural and horticultural crops. Conventional breeding programme is one of the option to develop resistant crop varieties but this practice is time consuming and non-durable because the resistance is quite often overcome by the development of new virulent virus strains. Further more, for fungally-transmitted viruses such as PMTV, no source of R gene have been reported in potato cultivars (R. M. Solomon-Blackburn, personal communication) and for others, resistance is not sufficiently strong to prevent infection when inoculum pressure is high or virus is delivered inside the susceptible hosts through zoospores. Transformation of plants, with segments of viral genome, frequently results in the development of plants that are resistant or immune to the virus from which the sequences were derived. This new approach called pathogen-derived resistance (PDR), will remain only a viable option for crop protection against virus infection. So far very little progress has been made to develop virus -resistant crop plant against furoviruses using PDR-strategies. This is because firstly the handling of the virus and vector is difficult and time consuming and secondly, most of the furoviruses are infecting cereals and transformation of such crops is difficult. However, there have been few attempts to obtain transgenic resistance to furoviruses, although it has been shown that beet protoplast expressing BNYVV CP are resistant to the virus (Kallerhoff et al., 1990). Sugar beet (Beta vulgaris L.) plants have been transformed with the BNYVV CP gene and challenged by bait tests with viruliferous soil. The trangenic plants exhibited decreased virus accumulation as compared with nontransgenic plants (Mannerlof et al., 1996). Extreme resistance or immunity has been established against PMTV using CP-gene of the virus in Nicotiana benthamiana. Three of transgenic lines were immune to infection following manual, graft or virus inoculation through viruliferous fungal zoospores (Reavy et al., 1994, 1995; Arif et al., 1999a). Virus challenge, through viruliferous zoospores into roots of the transgenic plants, can not overcome CR-MR (Reavy et al., 1995; Arif et al., 1999a). Recently, N. benthamiana lines were transformed using CP gene of PMTV (Scottish isolate) which conferred extreme resistance (93.5%) to 2 Swedish and 2 Danish PMTV isolates (Reavy et al., 1997). Development of resistance against 4 Scandinavian PMTV isolates, using CP-gene from Scottish isolate of PMTV, further indicated that CP-MR could confer resistance against isolates from different geographical areas. Nothing is known about durability of these transgenic resistance strategies in field. Mutation in RNA viruses might overcome such acquired resistance (Arif and Hassan, 2000). Ultimately, the question of durability and effectiveness can only be answered when transgewnic plants are put into large scale production over period of years. Durability of resistance could be improved by combing two or more genes conferring resistance (Beck et al., 1994) either transgene alone or a combination of transgenes and host genes (Barker et al., 1994a,b).
In future, it is hoped that genetic engineering of crop plants will provide effective protection against furoviruses. The strategies include the use of viral coat protein genes, the 3'-untranslated viral RNA sequence to compete with viral replicase, the viral nonstructural (e.g. replicase) polypeptides, catalytic RNA-cleaving ribozyme sequence, targeted to essential viral genes, defective RNA or protein species to block cell-to-cell spread, fungal acquisition or transmission of the furoviruses. The present paper reviews the available information on biological and molecular pathological properties of furoviruses, reported elsewhere in the world. Nothing is known about furoviruses in Pakistan and no work has been done so far on these viruses. S. subterranea, the causal agent of potato powdery scab disease and the vector of PMTV itself is a serious problem for potato production in Pakistan. PMTV have been suspected on the basis of characteristic symptoms but no detail studies have been done on this virus. SBWMV and P. graminis has also been suspected and symptoms of the virus have been observed in low lying areas of wheat fields in NWFP. The symptoms of PCV were commonly observed in groundnut in almost all groundnut growing areas and its identity has also been confirmed by ELISA, in selected plants (Delfosse et al., 1995). During survey of insect-transmitted viruses in sugar beet, the symptoms of BNYVV and BSBV were commonly observed in sugar beet growing of NWFP. Certainly many plant viruses, with fungal vectors, are infecting important crop plants and are potential threat to many cash and food crops in Pakistan. Detail and comprehensive scientific work, on fuoviruses, is extremely needed so that management practices can be devised to save agricultural crop against these viruses. This review will certainly serve as a base line to stimulate research on furoviruses and their fungal vectors in Pakistan and elsewhere in the world.
I express my sincere gratitude to many honourable members of International Working Group on Plant Viruses with Fungal Vector (IWGPVFV), who provided useful comments, suggestions, explanation and moral support, as well as unpublished results, pre-prints and reprints relevant to this review. With due apologies to any-one inadvertently omitted, special thanks go to Drs. B. Reavy, L. Torrance, H. Barker, T. M. A. Wilson, M. A. Mayo of Virology Department and J. M. Duncan and J. G. Harrison of Fungal and Bacterial Plant Pathology Department, Scottish Crop Research Institute, Invergowrie, Dundee, U. K. I am also grateful to Professor Dr. Sher Hassan and Professor Fida Mohammad Khan, NWFP Agricultural University, Peshawar, Pakistan, for their constructive comments on the text.