ABSTRACT
Primary structure of human granzyme H was aligned with cathepsin G and granzyme B. Besides, the 3D model of granzyme H was also constructed using the crystal co-ordinates of human cathepsin G. The active site of granzyme was predicted and mode of binding with substrate was discussed in the light of constructed model.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/pjbs.2001.110.111
URL: https://scialert.net/abstract/?doi=pjbs.2001.110.111
INTRODUCTION
Among the molecules proposed to be involved in cytotoxic T lymphocytes (CTL), Natural killer (NK) and lymphokines activated cell (LAK) mediated lysis are the granzymes, a family of serine protease stored in cytoplasmic granules of CTL, NK and LAK cells. These granzymes are essential for induction of target cell apoptosis (Yamashita et al.,1998). Jans et al. (1996) proposed that CTL and NK are able to kill their target cells through synergistic action of perforin and granzymes resulting in distinctive nuclear changes typical of apoptosis. It is reported by Haddad et al. (1991) that granzymes over expression is a marker of cytotoxic cell activation and should be further evaluated in patients with malignancies to delineate their potential value in predicting clinical outcome. In addition to the granzyme A and B, a third family member has been cloned in man and designed granzyme H. It was also reported that macular weight of granzyme H was approximately 30 K da and basic in nature. Granzyme H was isolated from lymph node of breast cancer patient (Khurshid, 2001). The intent of the present study is to construct 3D structure of Granzyme H isolated from lymph nodes of breast cancer patients.
MATERIALS AND METHODS
Amino acid sequences of granzyme B, H and cathepsin G were taken from Swiss protein data bank, Switzerland. A model of granzyme H was constructed using cathepsin G as template by program Modeler (Sanchez and Sali, 1997).
RESULTS AND DISCUSSION
Model of Granyme H: A rule based structural model of granzyme H was constructed on the basis of known 3D structure of other serine protease like cathepsin G using Modeler program. Cathepsin G was used as template . This shows 88.8% identity in 224 amino acid overlap. Among 224 residues of granzyme H, 223 carbon alpha atom can be superimposed on corresponding carbon alpha atom of cathepsin G.
Sequence and structure conservation among Cathepsin G and Granzyme H: Amino acid sequence of human granzyme H was compared with homologous human cathepsin G and granzyme B. Granzyme H is 88.8% identical with cathepsin G and 85% with granzyme B. (Fig. 1). It is reported by West et al. (1995) that granzyme H shows the highest degree of homology with granzyme B and cathepsin G and like these genes, consist of 5 axons separated to interons at equivalent position. It is therefore suggested that the ancestral gene of granzyme H is more likely related the cathepsin G and granzyme B than the other granzyme.
 | |
| Fig. 1: | Comparison of amino acid sequence of granzyme H with cathepsin G and granzyme B. Gaps introduced to optimize the alignment are denoted by dots. An asterisk indicates the conserved amino acid residue in all compared sequences. Arrows show amino acid ser11, asp181 and his 57 act as catalytic triad |
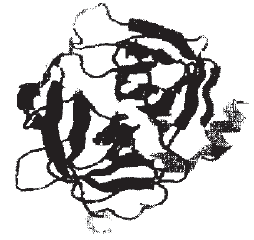 | |
| Fig. 2: | Ribbon diagram of granzyme H of polypeptide chain folded into 4α helixes (only in left domain) 913 sheets (in both domains) |
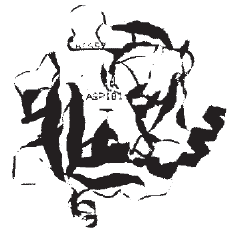 | |
| Fig. 3: | Ribbon plot of Granzyme H with catalytic sites Ser11, His57 and Asp181 |
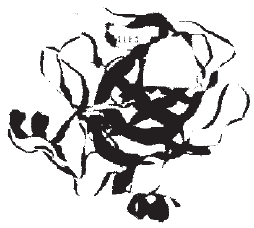 | |
| Fig 4: | Granzyme H with N-terminal lieu and C-terminal Lys |
Quality of model: Model consists of 1750 protein atoms consisting 145 hydrogen bonds. It was refined to a final crystallographic G factor of B0.10. Maximum deviation of model is 4.6. Ramachandaran plot of all main chain dihedral les shows that all non-glycemic residues are in favorable conformation with the exception of serine25. Which falls in more generously allowed region. Whereas leucine21 falls in disallowed region with phi-psi distribution of B0.29. 3D structure of enzyme was solved at 2.5 A resolution.
Tertiary fold: Structure of granzyme H is composed of 4 alpha helices and 9 beta strands with an overall α/β fold common for globular protein that can be classified as a slight variation of a doubly wound α/β domain. This reveals a conserved hydrophobic core corresponding to the central β strand and amphipathic α helices that form the interior of domain suggesting that the homologous granzyme H have a similar α/β fold. So this shows that structure is predominantly b structure with each of domain is a deformed b band. Domain A and B are approximately palindromic type. Overall tertiary folds are shown by ribbon representation in Fig. 2. N-terminal isoleucine and C-terminal lysine is stabilized by H bonding (Fig. 3). Connecting loops at N-terminal edge of β sheet surrounding the predicted active site region are significantly more conserved than loops at C-terminal edge of β sheet. This would be consistent with the importance of these loops both in substrate binding and in positioning the catalytic residues involved in hydrolysis reaction. So granzyme H sequence represents a functional gene expressed in activated T-cells. Stability of granzyme is enhanced by entropic effect of 3 disulfide bridges.
Active site: Side chain residue on the catalytic side are serine side chain hydroxyl methyl, histidine side chain imidazole and aspartate side chain. Hydrolysis of carboxyl methyl group and amide group catalyzed by granzyme H is expected to involve a catalytic mechanism similar to that of serine protease. In the catalytic domain of granzyme the 3 residues ser11, His57, asp181 are located in a cleft formed by loops at carboxyl ends of β strands (Fig. 4). Numerous H bonds stabilize the loops and side chains that form the active site. It is observed by Edwards et al. (1999) that eukaryotic serine protease have the order His, Asp, ser and subtilisin family is ordered Asp, His, Ser. Whereas granzyme H deviates from all of these having the order of His, Ser, Asp. It is noted that most hydrophobic amino acid is near to active center and these may protect the active center.
ACKNOWLEDGMENTS
Thanks are due to Dr M Jamil Qureshi, Chief Scientific Officer, Head Biological Chemistry Division, NIAB, Faisalabad for reviewing the article and giving fruitful suggestion.
REFERENCES
- Edwards, K.M., C.M. Kam, J.C. Powers and J.A. Trapani, 1999. The human cytotoxic T cell granule serine protease granzyme H has chymotrypsin-like (Chymase) activity and is taken up into cytoplasmic vesicles reminiscent of granzyme B-containing endosomes. J. Biol. Chem., 274: 30468-30473.
CrossRefDirect Link - Haddad, P., D. Jenne, J. Tschopp, M.V. Clement, D. Mathieu-Mahul and M. Sasportes, 1991. Structure and evolutionary origin of the human granzyme H gene. Int. Immunol., 3: 57-66.
CrossRefDirect Link - Jans, D.A., P. Jans, L.J. Briggs, V. Sutton and J.A. Trapani, 1996. Nuclear transport of granzyme B (Fragmentin-2) dependence on perforin in vivo and cytosolic factors in vitro. J. Biol. Chem., 271: 30781-30789.
CrossRefDirect Link - Khurshid, R., 2001. Serine protease granzyme H isolated from lymph nodes of breast cancer patient. J. Med. Sci., 1: 8-10.
CrossRefDirect Link - Sanchez, R. and A. Sali, 1997. Advances in comparative protein-structure modelling. Curr. Opin. Struct. Biol., 7: 206-214.
CrossRefPubMedDirect Link - Yamashita, Y., Y. Yatabe, T. Tsuzuki, A. Nakayama and Y. Hasegawa et al., 1998. Perforin and granzyme expression in cytotoxic T-cell lymphomas. Mod. Pathol., 11: 313-323.
PubMedDirect Link - West, A.H., E. Martinez-Hackert and A.M. Stock, 1995. Crystal structure of the catalytic domain of the chemotaxis receptor methylesterase, CheB. J. Mol. Biol., 250: 276-290.
CrossRefDirect Link








