Research Article
Conformation and Surface Properties of Gluten Treated with Protease from Germinating Wheat
Department of Biochemistry and Molecular Biology, University of Rajshahi, Rajshahi-6205, Bangladesh
In the previous paper, (Shaha et al., 1997 and Shaha and Hassan, 2001) it was reported that the functional properties of gluten including solubility, emulsifying and foaming properties, were greatly improved by treating with deamidase and protease from germinating wheat seeds. It was also suggested that the improved functional properties were mainly due to a deamidation process and not to a cleavage of peptide linkages in the gluten molecules. In addition, it was assumed that deamidated gluten by protease, improve its functional properties because this deamidation might induce conformational changes by increasing the electrostatic repulsion and decreasing the hydrogen bonding.
It has been recently reported that the surface structure of proteins plays an important role in the functional properties of food protein. Some investigators (Naksi et al., 1980; Naksi, 1980; Matsudomi et al., 1986 and Kato et al., 1983) have reported that surface hydrophobicity showed a significant correlation with the emulsifying properties of the various proteins. Therefore, it can be presumed that the functional properties improved by deamidation may be due to an increase in the effective hydrophobicity of gluten. However, little is known about the surface structure of gluten. Therefore, further investigation on the surface structure of gluten needed to be done to correlate protein functionality with conformation. This paper describes the effect of conformational changes on the surface properties of deamidated gluten by crude protease from germinating wheat seeds.
Materials and Methods
Source and preparation of gluten: Wheat flour was collected from Local Shahab Bazer Market, Rajshahi (Bosundhara Flour Milling Ltd., Dhaka, Bangladesh). Insoluble wheat protein complex, gluten, was prepared from a flour dough washed until the washes were free from soluble protein. The gluten ball thus obtained was dialyzed against distilled water and then freeze-dried.
Extraction and purification of protease from germinating wheat seeds: Germinated wheat seeds at 72 hours were taken in a mortar, ground and homogenized uniformly with pre-cooled distilled water in a ratio 4:50 (w/v). The water extract was then occasionally stirred with a glass rod and kept over night at 4°C. The clear filtrate was collected by cooling and centrifuging at 6000 rpm at 10 min. The supernatant was adjusted to 80% saturation by adding ammonium sulfate solution. The precipitate was dissolved in minimum volume of pre cooled 5 mM phosphate buffer (pH 7.4) and dialyzed against 5 mM phosphate buffer (pH 7.9) for 24 h in a cold chamber at 10°C. After centrifugation, the clear supernatant was used as crude enzyme extract. The crude enzyme extract was then loaded on Sephadex G-75 and then on DEAE-Cellulose column for further purification. Both columns were washed with 5 mM phosphate buffer pH 8.0, containing 1 M sodium chloride. Enzymatically active protein fractions were collected and then freeze dried (Shaha et al., 2002).
Preparation of protease treated gluten samples: Hundred milliliters of a 2% gluten suspension was prepared by phosphate-citrate buffer pH 6.9 and then 100 mg of partially purified protease (Shaha et al., 2002) was added to the protein solution. The mixture was incubated at 31°C for 2 h with shaking. After incubation, the reaction mixture was immediately heated for 30 sec at 100°C to inactivate the protease. The enzymatically treated gluten solution thus obtained was centrifuged and the clear solution was then dialyzed against distilled water exhaustively and freeze-dried.
Chemical analysis: From the protease treated gluten (protease from germinating wheat seeds) sample amide nitrogen was determined by using glutamate dehydrogenase according to the method of Kun & Kearney (1974). Protein concentration was determined by the method of Lowry et al. (1951), using bovine serum albumin as a standard. SH-groups and SS-groups in the gluten protein were determined according to the method of Beveridge et al.(1974) using Eliman’s reagent.
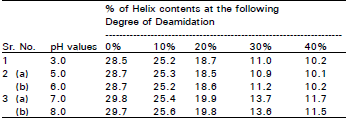
Circular dichromism (CD) analysis: Thirty mg of the protease treated gluten sample was dispersed in 10 ml of 0.1 M Mcllvaine buffer, at pH 3.0, 7.0 and 8.0. Each of these gluten suspensions was homogenized lightly in an Ultra Turrax Disperser TP 18 (Hansen & Co. Ltd.) and then filtered through filter paper (Toyo Roshi. No. 5B). The soluble protein solution thus obtained was used for CD analysis and the determination of protein concentration. Circular dichromism was measured on a Jasco Model J-20 C spectro-polarimeter under the far-ultraviolet region condition (200 ~ 300 nm) with a 0.01 cm path length of cell at 25°C. The helix contents were calculated from the mean residue elliptically at 222 nm, using 110 as the residue weight of proteins. At pH 7.0 and pH 8.0, all of the gluten samples had a similar value for helix contents but at pH 3.0 the value for helix content slightly less than other pH values.
Determination of surface hydrophobicity: Thirty mg of the protease treated gluten sample was dispersed in 10 ml of 0.01 M phosphate buffer, pH 7.0. The soluble protein solution was obtained in a similar manner to that described in CD analysis. This solution was used for the determination of surface hydrophobicity, which was determined by the fluorescence technique using cis-parinaric acid (Kato et al., 1980). The initial slope (So), i.e. fluorescence intensity 1% protein, was calculated from plots of the fluorescence intensity vs. protein concentration.
Measurement of surface properties: Surface tension was measured with a 0.2% protein solution in a 0.01 M phosphate buffer (pH 7.4) on a Dunojiy Interfacial Tensiometer (Kyowa Kagaku K.K., Japan) at 25°C.
The emulsifying activity index was determined according to the method of Pearce and Kinsella (1978). To prepare the emulsions, 1.8 ml of corn oil and 4.2 ml of 0.2% protein solution in a 0.01 M phosphate buffer (pH 7.4) were homogenized in disperse at 12000 rpm for 1 min at 25°C. Emulsion stability was calculated from the following procedure. Aliquots (100 μl) of the emulsion obtained above were taken from the bottom of the container after different time intervals (0,1,2,10 and 30 min) and diluted emulsion was then determined at a wave length of 500 nm and indicated as a turbidity. The turbidity half time was calculated from plots of turbidity vs time-course. The emulsion stability was expressed as the value of half-time x initial turbidity.
Conformational changes by the protease treated gluten: In the previous papers (Shaha et al., 1997; Shaha and Hassan, 2001) it was suggested that deamidation of gluten by deamidase (less than 40%) from germinating wheat seeds may induce conformational changes without cleavage of the peptide linkages in the gluten molecules. On the other hand, conformational changes due to SS disruption may occur during the deamidase reaction of gluten. Therefore, the contents of the SH-groups and SS- groups in the protease treated gluten were measured (Fig. 1). The relationship between the surface hydrophobicity and the degree of deamidation in gluten is shown (Fig. 2). Surface hydrophobicity increased linearly in proportion to the degree of deamidation and a close correlation (r=0.97, p< 0.01) was observed between these two parameters.
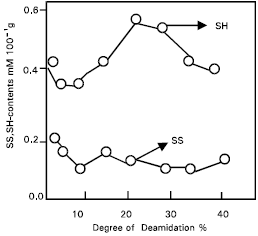 | |
| Fig. 1: | Changes in SH-and SS-contents during deamidation of protease treated. |
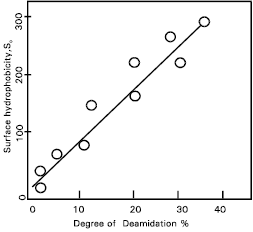 | |
| Fig. 2: | Relationship between surface hydrophobicity and deamidation of protease treated gluten. |
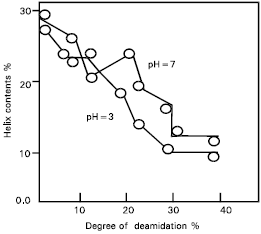 | |
| Fig. 3: | Relationship between helix contents and deamidation of protease treated gluten. |
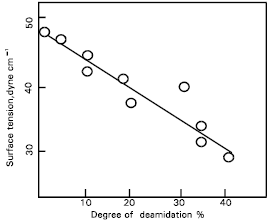 | |
| Fig. 4: | Relationship between surface tension and deamidation of protease treated gluten. |
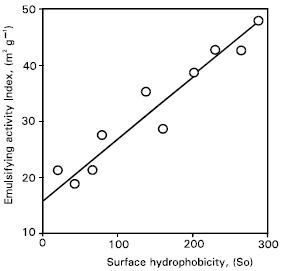 | |
| Fig. 5: | Relationship between emulsifying activity index and surface hydrophobicity of protease treated gluten. |
The relationship between helix content and the degree of deamidation in the gluten is also presented (Fig. 3). The helix contents decreased curvilinearly with the increase in deamidation. The relationship between surface tension and the degree of deamidation of protease treated gluten is shown (Fig. 4).
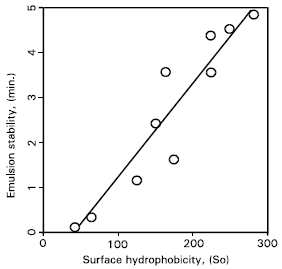 | |
| Fig. 6: | Relationship between emulsion stability and surface hydrophobicity of protease treated gluten. |
 | |
| Fig. 7: | Relationship between surface tension and surface hydrophobicity of protease treated gluten. |
| Table 1: | Relationship between Helix contents and deamidation of protease treated gluten at different pHs |
 | |
Relationship between surface hydrophobicity and surface properties: We have already reported that the surface properties of gluten were improved markedly by deamidation. It is probable that the main structural factor governing the functional properties of protein may be the hydrophobic regions at the protein surface which enhance the interaction between the protein and other molecules, such as lipids. Therefore, the surface properties of deamidated gluten were correlated with the surface hydrophobicity.
The relationships of emulsifying activity and emulsion stability with surface hydrophobicity of the deamidated gluten are given (Figs.5, 6). The emulsifying activity index correlated linearly with surface hydrophobicity and the correlation coefficient was 0.88 and highly significant (p< 0.01). The emulsion stability also correlated linearly with surface hydrophobicity and the correlation coefficient was 0.96 and highly significant (p < 0.01). There was a linear and –ve relationship between the surface tension and surface hydrophobicity of deamidated gluten (Fig. 7).
Discussion
Changes in the SH and SS groups respectively from the protease treated gluten were non-significant, although the possibility of conformational changes due to the SS interchange reaction still exists. In addition, the helix content of the protease treated gluten was measured to elucidate these conformational changes (Fig. 1). Results suggest that the hydrophobic regions are exposed at the molecular surface in proportion to the conformational changes caused by deamidation (Fig. 2). The conformational changes (Fig. 3) may be caused mainly by the increased electrostatic repulsion and the decreased hydrogen bonding as a result of deamidation. When the degree of deamidation in gluten become about 40%, the helix contents of the gluten decreased to about 10 from 30% of native gluten. At pH 7.0 and 8.0, all protease treated gluten samples had similar value for helix contents but at pH 3.0 the value for helix contents was slightly less than other pH values. However, the helix contents were always retained at about 10% even when the degree of deamidation was over 40%. From the results of relationship between surface tension and the degree of deamidation (Fig. 4), it was suggested that a stable structure may partially exist in the gluten molecules. Surface tension decreased linearly in proportion to the deamidation and the correlation coefficient between these parameters was –0.90 and highly significant (p< 0.01). So, the deamidation of gluten caused a pronounced reduction in surface free energy at the interface between air and water, probably due to induction of an amphophilic nature by the remarkable increase in surface hydrophobicity.
Results about the emulsifying activity and emulsion stability (Fig. 5 & 6), confirmed that the emulsifying properties of gluten were improved greatly by deamidation, correlating linearly with increased surface hydrophobicity. A good correlation (r=–0.90, p=0.01) was observed between the surface tension and surface hydrophobicity of deamidated gluten. Thus, it was elucidated that the surface properties of deamidated gluten were improved, depending mainly on the surface hydrophobicity.
In conclusion, a marked increase in surface hydrophobicity is the main structural factor for an improvement in functional properties by the deamidation of gluten. Induction of an amphophilic nature, due to an increase in surface hydrophobicity, caused a decrease in free energy at the surface of the deamidated gluten and endowed good surface properties.
Some of the analysis was helped by Prof. A. Kato, Laboratory of Biological Science, Yamaguchi University, Japan. This work was done in the Department of Biochemistry & Molecular Biology, Rajshahi University, Rajshahi, Bangladesh and was supported by the grant-in-aid (Grant-1-99-335) from Ministry of Science and Technology, Dhaka, Bangladesh.