Research Article
Glutamine Synthetase in Harvested Asparagus Spears:Changes in Activity and Gene Expression During Storage
Not Available
Toshiyuki Matsui
Not Available
Haruo Suzuki
Not Available
Yusuke Kosugi
Not Available
Harvested asparagus (Asparagus officinalis L.) spears accumulate ammonia in their tips towards the end of shelf-life (Hurst et al., 1993). Glutamine synthetase (GS; EC 6.3.1.2) is the primary enzyme responsible for assimilating ammonia in plants (Joy, 1988). GS catalyses the ATP-dependent conversion of glutamate to glutamine utilizing ammonia as a substrate. Treatment of asparagus with phosphinothricin (PPT), a potent inhibitor of GS in plants, results in a rapid accumulation of ammonia in the spear tips and a reduction of shelf-life (Hurst et al., 1993). Ammonia accumulation is thought to be a factor contributing to the perishability of harvested asparagus spears and occur due to alterations in GS activity after harvest. As ammonia is thought to be toxic to plant cells, the changes in ammonia content and GS gene expression in both top and bottom portions of asparagus spears were studied in order to further understand the factors influencing ammonia assimilation during storage.
Asparagus spears like leaf cells of some other higher plant species, contain two isoforms of GS that are distinguished by their subcellular position: GS-1 is only present in the cytosol, whereas GS-2 is exclusively detectable in the chloroplast matrix (Downs et al., 1994). Little is known about the regulation of GS gene expression in plants or the significance of multiple GS family members. Expression of some of these members is localized to the phloem elements (Brugière et al., 2000), the N2-fixing root nodules of legumes (Edwards et al., 1990), cotyledons (Cantón et al., 1999) and senescing leaves (Bernhard and Matile, 1994; Brugière et al., 2000; Masclaux et al., 2000), suggesting a role in N uptake, translocation and mobilization. Although both the GS-1 and GS-2 cDNA clones have become available for many mono- and dicotyledonous species, for asparagus they have not been isolated yet. In the EMBL/GenBank/DDBJ databases only one cDNA clone (“pTIP 23; X71361) was found submitted as Asparagus officinalis mRNA for glutamine synthetase.
In the study one partial cDNA clone was isolated and sequenced that was identical to the “pTIP 23 in the databases and was used as a probe for northern blot analysis of total GS without separation of cytosolic and plastid forms.
Materials and Methods
Plant material: Asparagus spears were harvested from farmer’s field in Kagawa Prefecture, Japan during June-July, 2001. Spears were immediately brought to the laboratory (Faculty of Agriculture, Kagawa University, Japan) and stored at 200C in perforated plastic bags for up to four days in darkness. After storage treatment spears were cut into two equal halves (designated as the top and bottom portion), weighed and immediately stored at –800C until needed for GS and ammonia assay and for total RNA extraction.
Enzyme extraction: Approximately 5 g sample from each portion of the spear were homogenized under ice cold condition (ca. 0-4°C) with 1% polyvinylpolypyrrolidone (PVPP), proportional to the sample weight and 1 g of sea sand in Buffer A by using a mortar and pestle. One ml extraction buffer per gram fresh weight of plant materials was used. Extraction was performed according to the method of Hurst and Clark (1993), in which Buffer A contained 50 mM Tris-Hcl (pH 7.6), 10 mM MgSO4, 1 mM EDTA, 1 mM dithiothreitol (DTT), 12 mM 2-mercaptoethanol, 5 mM glutamate and 100 ml/liter glycerol. The homogenate was squeezed through a four-layer cotton cloth and the filtrate was centrifuged at 11 000 x g for 10 min. The residual tissues were re-extracted in 5 ml Buffer A and dialyzed with 40 times dilution of the same buffer for 1 h and then centrifuged. The resulting supernatant were mixed together and used for the enzyme assay.
GS and ammonia assay: GS was measured in a total volume of 1.0 ml. The assay mixture contained 80 mM L-glutamate-Na, 500 mM tricine-KOH Buffer (pH 7.0), 600 mM NH2OH, 200 mM MgSO47H2O, 10 mM diethylenetriamine pentaacetic acid (DTPA), 80 mM ATP and 800 mM mercaptoethanol. After incubating at 35 °C for 8 min, the reaction was stopped by adding 1 ml ferric reagent (25 ml FeCl3 6H2O, 50 ml HCl and 20 ml TCA). GS activity was measured using a double beam spectrophotometer (Shimadzu model UV-150-02) at 540 nm.
For assay of ammonia 5 g samples from each portion were extracted with 10% Trichloroacetic acid and centrifuged at 11 000 x g for 10 min as described by Kun and Kearney (1974). Ammonia content of the supernatant was determined from a triplicate 500 μl sample by adding 200 μl 0.5 M tris-buffer (pH 8.0), 100 μl 0.1 M 2-oxoglutarate solution (pH 7.4), 30 μl 8 mM β–NADH solution, and 150 μl distilled water. The absorbance was recorded at 340 nm against a reagent blank.
RNA isolation: Total RNA was extracted according to the Hot Borate method of Wan and Wilkins (1994).
Amplification of poly (A)+ RNA by RT-PCR: The first strand cDNA was synthesized from 2 g of the total RNA by reverse transcriptase with Oligo-(dT) primer according to the instruction of SUPER SCRIPTTM Preamplification System for First Strand cDNA Synthesis (GIBCOBRL, Tokyo, Japan). PCR was performed in a total volume of 25 μl containing the first strand cDNA reaction products, 10 x PCR Buffer, MgCl2, dNTP, First Start Taq DNA Polymerase (Roche) and primers. The primers (5’-TCNAGCACNGGGCARGCNCC -3’ as the sense primer and 5’- TCRTTNCCYTCNCCRTANGC -3’ as the antisense primer) were designed and synthesized on the basis of amino acid domains conserved in various GS genes. The Sal 1 and Not1 restriction site sequences were also included at 5’-end of the sense and antisense primer, to facilitate cloning of PCR product. The PCR procedure started with 10 min at 950C and was carried out 35 cycles of 30 s at 95, 50 and 720C, and 10 min at 720C with ASTEC Program Temperature Control System PC-700. The PCR products were confirmed by agarose gel electrophoresis.
Cloning and sequencing of cDNA: The amplified cDNA was ligated to the plasmid pSPORT1 and cloned into Escherichia coli (DH-5α ) Not 1- Sal 1 – cut (BRL, Tokyo, Japan). Sequencing was performed by the cycle sequencing method using GATCR-Bio Cycle sequencing Kit and a DNA sequencer GATC 1500 Long-Run system (GATC Gmbh, Konstanz, Germany).
Sequence data analysis: Sequence analysis was performed using computer software GENETYX-MAC Ver.7. Homology searches with the Genbank and the EMBL databases were performed using the homology program in the software.
Preparation of the digoxigenin (DIG)-UTP-labeled RNA probe: The cloned RT-PCR product including the encoded region of GS gene was cleaved by Not 1 and Sal 1 from the pSPORT1 vector that had been amplified in Escherichia coli (DH-5α) and it was purified and recovered by gel electrophoresis. AntisenseDIG-labeled RNA probe was prepared according to the instructions of DIG RNA Labeling Kit (Boehringer Mannheim) using SP6 RNA polymerase.
Northern blot analysis: Ten μg of total RNA was subjected to electrophoresis on a 1.0% agarose (Type II) gel containing 20 x MOPS and 37% formaldehyde. After electrophoresis for 30 min, RNA was visualized with ethidium bromide under UV light to confirm equal loading of RNA in each lane. RNA was transferred to a positively charged nylon membrane HybondTM- N+ (Amersham Pharmacia Biotech) by capillary action with 20 x SSC and then after drying the membrane RNA was fixed under UV. The membrane was prehybridized at 500C with 5 x SSPE, 5 x Denhart’s solution, formamide and 10% SDS for 3 h. Hybridization was performed at 500C using the gene specific antisense DIG-labeled RNA probe for 24 hours using the same prehybridization buffer. After hybridization, the membrane was washed twice with 2 x SSPE containing 0.1% SDS for 10 min at room temperature, once with 1 x SSPE containing 0.05% SDS for 15 min at 650C and once with 0.2 x SSPE for 10 min containing 0.05% SDS at 650C. The membrane was also washed with Buffer A containing maleic acid and tween 20 at room temperature and was blocked with 2% blocking reagent in maleic acid buffer for 30 minutes. Subsequently, the membrane was incubated with anti-digoxygenin –AP, fab fragments (Boehringer Mannheim) in the blocking buffer for 30 min. Signals were detected by color reaction using 5-bromo-4-chloro-3-indolyl- phosphate and 4-nitro blue tetrazolium chloride as the substrate.
Results
Ammonia: A general increase in ammonia content for both top and bottom portions of the spears was observed after harvest (Fig. 3). Ammonia accumulation increased to about 40% of initial level after 72 h storage and reached about 60% of the initial level at the end of 96 h storage period. Higher (almost double) accumulation of ammonia was found in the top portion than in the bottom portion of the spears.
GS activity: GS activity began to decline at harvest and continued throughout the storage period in both top and bottom portions (Fig. 1A). After 96 h storage period the activity in the top and bottom portions declined to about 22 and 24% of the initial level respectively. The GS activity in the top portion was significantly higher than the bottom portion of the spears.
GS gene expression: cDNA for GS amplified by RT-PCR which corresponded to about 695 bp was cloned and sequenced. The 695 bp cDNA for GS was identical (97.6 %) to the same region of the cDNA (Fig. 2) that encoded Asparagus officinalis m RNA for GS (pTIP23; X71361). Therefore, the 695 bp cDNA clone was used as a probe for northern blot analysis to determine the levels of transcripts for GS.
The levels of transcripts for GS decreased after harvest in both top and bottom portions of the spears (Fig. 1B) and correlated well with enzyme activity. Expression of GS gene in the top portion was clearly observed up to 24 h of storage and remained detectable after 96 h storage period. But the expression in bottom portion was barely detectable that may reflect in the much lower GS activity in that portion.
Ammonia is toxic to plant cells at high concentrations and is normally assimilated if produced within the plant (Miflin and Lea, 1980). The accumulation of ammonia in senescing leaves has been shown to coincide with almost complete disappearance of GS (Peters and Van Laere, 1992).
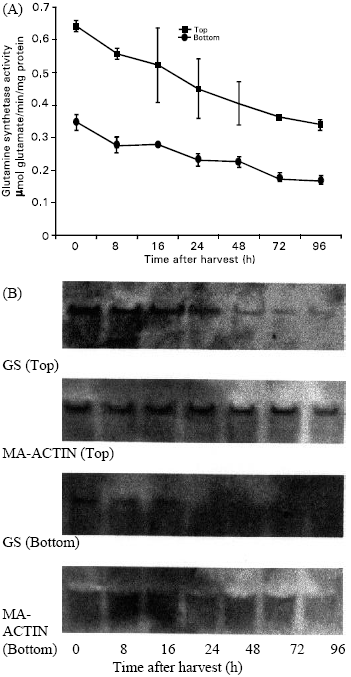 | |
| Fig. 1: | Time courses of GS activity and gene expression in the top an bottom portions of the spears held at 20 °C (A) Changes in GS activity. Vertical bars indicate SE. (B) Northern blot analysis of total RNA (10 μg per lane) showing gene expression. MA-Actin was used as internal control. |
In this study the same trend in which, GS activity and the level of transcripts for total GS declined in both top and bottom portions of the spears was observed with an increased in ammonia accumulation to about 60% after 96 h storage period (Fig. 1B). Hurst and Clark (1993) reported 30% decline of total GS activity in harvested spear tips after 5 day storage at 20°C. The levels of plastid GS activity, polypeptide and mRNA all declined in the tips of harvested asparagus spears (Downs et al., 1994).
 | |
 | |
| Fig. 2: | Comparison of synthesized GS probe sequence with asparagus GS nucleotide sequence (pTIP23) in database (1st line: Synthesized GS prob; sequence size: 695 and 2nd line: Asparagus GS nucleotide in the database; sequence size: 1651). Asterisks (*) denote the nucleotide residues those are identical (97.6%/695bp) |
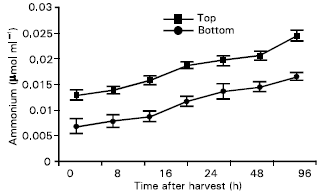 | |
| Fig. 3: | Ammonia content in the top and bottom portions of the spears. Vertical bars indicate standard error |
It has been reported that plastid GS has a role in assimilating ammonia produced during photo respiration in higher plants (Edwards and Coruzzi, 1989). Hurts et al. (1993) reported that although about 65% of tip GS activity was lost when spears were held at 20°C ammonia accumulation was not due to limiting GS levels. They have studied postharvest inhibition of glutamine synthetase and used PPT 40 and 200 ppm which reduced GS activity by 50 and 80%, respectively, but did not increase ammonia accumulation. Ammonia accumulation was only enhanced, when greater than 95% of GS was inhibited. Thus, it seems that there is a critical level of GS activity necessary for the postharvest life of asparagus, and that under normal postharvest conditions, this critical level is exceeded and is more than the requirement of ammonia salvage. It is likely that ammonia accumulation results from a shortage of carbon skeletons required for its assimilation, i.e. the demand for respiratory carbon ultimately outstrips the availability of carbon for ammonia reassimilation, and ammonia accumulation ensues.
In conclusion, the increase in ammonia content and decrease in total GS activity in both top and bottom portions of the spears is a response to significant stress after harvest. Perhaps as a consequence of general metabolic decline occurring at this time (King et al., 1993) or may be the consequence of autophagic process (Journet et al., 1986) induced by carbohydrate deprivation.
Acknowledgments
The financial support of the Ministry of Education, Science, Sports and Culture of Japan under scholarship program for foreign students is gratefully acknowledged.