Research Article
Isolation and Characterization of Three CDNAs Encoding CHH-family Peptides from the Crab Oziotelphusa senex senex
Shri Venkateshwara University, India
P. Sreenivasula Reddy
Shri Venkateshwara University, India
An important part of the neuroendocrine system in decapod crustacean is situated in the optic ganglia of the eyestalk and consists of a cluster of 150-200 peptidergic neurons form the X-organ in the medulla terminalis ganglion. The enlarged axon terminals fuse and form sinus gland, a neurohaemal organ where the secreted products are stored. Several large peptide hormones are synthesized and released from the X-organ-sinus gland complex which includes crustacean hyperglycemic hormone (CHH; regulates carbohydrate metabolism), molt-inhibiting hormone (MIH; inhibits ecdysteroid synthesis by the Y-organ), vitellogenesis-inhibiting hormones (VIH; inhibits the onset of vitellogenesis) and mandibular organ-inhibiting hormone (MOIH; inhibits methyl farnesoate synthesis from mandibular organ) (Reddy and Ramamurthi, 1999; Huberman, 2000). Among these, more than 75% of total peptide hormone produced belongs to hyperglycemic hormone. All these hormones are structurally related and some exhibit overlapping biological activities thereby form a peptide family referred to as the crustacean hyperglycemic hormone family (Keller, 1992).
In recent years, more attention is paid to characterize CHH-family peptides due to their involvement in the regulation of growth and reproduction in crustaceans. The CHH-family peptides were isolated and characterized from many crustacean species (Reddy and Ramamurthi, 1999; Huberman, 2000; Chang, 2001). The CHH-family peptide genes were isolated from the prawns Fennropenaeus chinensis (Wang et al., 2003), Macrobrachium resenbergii (Yang and Ranga Rao, 2001; Chen et al., 2003), Metapenaeus ensis (Gu and Chan, 1998a; 1998b; Gu et al., 2000), Penaeus japonicus (Ohira et al., 1997a and b, 2005), Penaeus monodon (Davey et al., 2000; Udomkit et al., 2000; 2004) and Penaeus vannamei (Sun, 1994, 1995), the lobsters Homarus americanus (Chang et al., 1990; Reddy et al., 1997; De Kleijn and Van Herp, 1998; De Kleijn et al., 1998) and Nephrops norvegicus (Edomi et al., 2003), the crayfishes Orconectes limosus (De Kleijn et al., 1994) and Procambarus bouvieri (Aguilar-Gaytan et al., 1997) and the crabs Callinectes sapidus (Lee et al., 1995), Cancer magister (Umphery et al., 1998), C. pagurus (Tang et al., 1999; Lu et al., 2000; 2001), Carcinus maenas (Weidemann et al., 1989; Gromoll and Weidemann, 1990), Libinia emarginata (Liu et al., 1997) and Uca pugilator (Klein et al., 1993). The data revealed polymorphism of the CHH neuropeptides in several decapod crustacean species. CHH isoforms were found in the crayfish Orconectes limosus (Kegel et al., 1991), Procambarus bouvieri (Huberman and Aguilar, 1988) and in the lobster Homarus americanus (Tensen et al., 1989). Multiple copies of these genes with minor variations were also reported from different species.
In the present study three cDNAs encoding CHH-family peptides from eyestalk neural tissue of the crab Oziotelphusa senex senex were isolated and sequenced. The divergence of deduced amino acid sequences were analyzed and compared with crab CHH-family peptides.
MATERIALS AND METHODS
Procurement of animals and housing: Intact, inter molt (Stage C4) adult freshwater crabs, Oziotelphusa senex senex Fabricius with a body weight of 30±3 g and body size of 39±3 mm were collected from the rice fields and irrigation canals around Tirupati, Andhra Pradesh, India. The crabs were maintained in animal facility, department of Biotechnology, Sri Venkateswara University, Tirupati. Animals were housed 6-8 per glass aquaria (length: width: height = 60:30:30 cm) with sufficient ambient medium (salinity: 0.5 ppt) and transferred to fresh medium everyday. They were acclimatized to the laboratory conditions (temperature 27±1°C; relative humidity 75% and a light period of 12 h) for 10 days before being used for experimentation. During their sojourn, the crabs were fed with sheep meat ad libitum. All the experiments were conducted in department of Biotechnology, Sri Venkateswara University, Tirupati.
Tissue collection and RNA isolation: The eyestalks from the crab Oziotelphusa senex senex were collected immediately after ice anesthetization. The eyestalks were stored at -196°C (in liquid nitrogen) until further use. A total of 20 eyestalks per reaction were taken out from the liquid nitrogen, the neural tissues were dissected out and used for total RNA isolation. The total RNA was isolated using the Perfect RNATM Eukaryotic, Mini Purification kit (Eppendorf, Hamburg, Germany) and the isolation was carried out according to the manufacturer’s protocol.
Primer designing: Primers were designed based on the conserved amino acid sequences of the CHH-family peptides from different crustacean species (Reddy and Ramamurthi, 1999; Lacombe et al., 1999; Chen et al., 2005). The highly conserved regions are between 20 and 30 amino acids with only one or two amino acid variations.
| Table 1: | Primers designed for PCR and their annealing temperatures |
 | |
| FW: forward primer; REV: reverse primer | |
These regions were decoded into oligonucleotides using codon preference table which includes highly accepted codon for a particular amino acid. Table 1 shows the primers designed for PCR amplification with annealing temperatures. All the designed forward primers were synthesized by Genetix, Hyderabad (made in Operon) consists of 17 nucleotides and the reverse primer used in the present study is Oligo (dT)18 (Ferments).
One step RT-PCR and cloning of amplified products: The first strand cDNA was synthesized with 500 pg of total RNA, followed by the polymerase chain reaction with the kit cMasterTM RTplus PCR System and cMasterTM RT (Eppendorf, Hamburg, Germany) according to the manufacturer’s protocol. Individual reactions were carried out with each forward primer. Each amplified product was cloned into PQE30 UA vector (QIAexpress UA Cloning Kit; QIAGEN, Valencia, USA) separately. The cloning was performed according to the manufacturer’s protocol. The recombinant plasmid was transformed immediately into the E. coli strain JM 109 (Genei, Bangalore, India) by the method of Cohen et al., (1972). The transformed cells were screened by spreading the cells (100 μL) on ampicillin (100 μg mL-1) containing M9 minimal agar medium plates spread with 40 μL of 2% X-gal (in dimethyl formamide) and 40 μL of 20% Isopropyl thiogalactoside (IPTG, in sterile distilled water). The bacterial cells were allowed to grow for 12 h at 37°C. The white colonies were isolated and sub-cultured in ampicillin (100 μg mL-1) containing M9 minimal medium. The culture is used for isolation of recombinant plasmid DNA.
Isolation of plasmid DNA and sequencing: The plasmid was isolated using GeNeiTM Ultrapure Plasmid Midi Prep Kit (Genei, Bangalore, India) according to the manufacturer’s protocol. Each clone was grown (12 h) in a separate pre sterilized conical flask containing ampicillin (100 μg mL-1) M9 minimal medium (100 mL) by inoculating single transformed colony. The quality of isolated plasmid was checked on 1% agarose gels. The insert size was confirmed by digesting the recombinant plasmid with 40 units of EcoR V and Sac1 for 2 h at 37°C and nucleotide sequence was determined for all the positive clones using the sequence facility at MWG Biotech Private Limited, Bangalore, India.
Analysis of sequenced data: The nucleotide sequences were aligned against the existing nucleotide sequences of the DNA data Bank at NCBI using BLASTN (Basic Local Alignment Search Tool) version 2.2.12 with expect value 10 and word size 11. The same nucleotide sequences were also searched against the existing protein sequences of the data bases at NCBI using BLASTX version 2.2.12 with expect value 10 and word size 3. The translation of coding sequences was done using win gene version 2.31. The amino acid sequences were aligned using CLUSTALX version 1.81. The obtained alignments were viewed and presented graphically using BioEdit version 5.0.9 and the phylogenetic tree was constructed and analyzed.
After cloning the amplified PCR products, three clones containing the expected insert size were obtained in the present study using the designed primers. Out of three PCR products (Oss-SGP1, Oss-SGP2 and Oss-SGP3), two obtained with the forward primer 3 (FW 3: 5’ TGCGAGGACTGCACCAA 3’) and one with forward primer 4 (FW 4: 5’ TGTGAGGATTGCTACAA 3’). Using primers 1, 2 and 5 did not result in amplification of DNA in polymerase chain reaction. The sizes of amplified DNAs were between 336-418 base pairs. The open reading frames of these cDNAs are more conserved (>75%) when compared with 3’ non-coding regions (<60%). All the clones have a polyadenylated site (AATAAA) near the poly(A) tail. The sequence of these cDNAs were determined and the deduced amino acid sequences were translated (Fig. 1-3).
 | |
| Fig. 1: | Nucleotide and deduced amino acid sequence of cDNA encoding CHH family peptide in Oziotelphusa senex senex (Oss-SGP 1). The deduced amino acid sequence is shown below the nucleotide sequence. Asterisk mark the termination codon. The putative polyadenylation signal is underlined. Arrows represent forward and reverse primers |
 | |
| Fig. 2: | Nucleotide and deduced amino acid sequence of cDNA encoding CHH family peptide in Oziotelphusa senex senex (Oss-SGP 2). The deduced amino acid sequence is shown below the nucleotide sequence. Asterisk mark the termination codon. The putative polyadenylation signal is underlined. Arrows represent forward and reverse primers |
 | |
| Fig. 3: | Nucleotide and deduced amino acid sequence of cDNA encoding CHH family peptide in Oziotelphusa senex senex (Oss-SGP 3). The deduced amino acid sequence is shown below the nucleotide sequence. Asterisk mark the termination codon. The putative polyadenylation signal is underlined. Arrows represent forward and reverse primers |
The deduced amino acid sequences of Oss-SGP 1-3 were aligned with other published sequence of CHH-family peptide genes of different crab species indicates (Fig. 4) that Oziotelphusa clones shows the highest amino acid identity to that of the CHHs of Pachygrapsus marmoratus (54.0%) and Scylla serrata (47.0%). However, the derived amino acid sequences widely diverged with the MIHs of Carcinus maenas, Cancer pagurus, Scylla serrata and Cancer magister (65.0-67.0%) and with MOIH of Cancer pagurus (73.0%). The result suggests that they may arise from individual polymorphism.
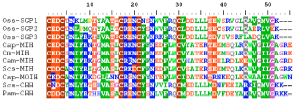 | |
| Fig. 4: | CLUSTALX (1.81) formatted amino acid alignment of partial mature sinus gland peptides from the crab Oziotelphusa senex senex (Oss-SGP 1-3), Cancer pagurus (Cap-MIH; Lu et al., 2001; GenBank accession No. AJ245380, Cap-MOIH; Tang et al., 1999; GenBank accession No. AJ245378), Carcinus maenas (Cm-MIH; Klein et al., 1993; GenBank accession No.X75995), Scylla serrata (Scs-CHH; Tsai and Lee, 2003; GenBank accession No. AY372181, Scs-MIH; Qui and Zhang 2002; GenBank accession No. AY083797), Cancer magister (Cam-MIH; Umphery et al., 1998; GenBank accession No. AF031493) and Pachygrapsus marmoratus (Pam-CHH; Toullec and Spanings-Pierrot, 2002; GenBank accession No. AY094983). The identified sequences among all the peptides are highlighted |
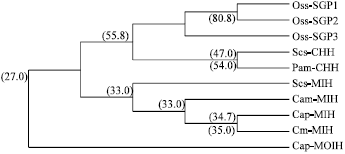 | |
| Fig. 5: | The dendrogram created by PileUp with the partial sequence of Oziotelphusa senex senex (Oss-SGP 1-3), Cancer pagurus (Cap-MIH; Lu et al., 2001, Cap-MOIH; Tang et al., 1999), Carcinus maenas (Cm-MIH; Klein et al., 1993), Scylla serrata (Scs-CHH; Tsai and Lee, 2003, Scs-MIH; Qui and Zhang, 2002), Cancer magister (Cam-MIH; Umphery et al., 1998) and Pachygrapsus marmoratus (Pam-CHH; Toullec and Spanings-Pierrot, 2002). The homology percentage with Oss-SGP 1 was mentioned in the parentheses |
To gain an understanding of the evolutionary relatedness of Oziotelphusa eyestalk sinus gland peptides with other known sequences of CHH-family neuropeptides, the phylogenetic analysis was carried out (Fig. 5). The phylogenetic analysis is similar to the analysis of deduced amino acid sequences. The phylogenetic tree shows that the crab sinus gland peptides (SGP-1 to 3) are structurally more similar to CHHs (<54%) than MIHs (<35%) and MOIH (27%).
In the present study the total RNA was isolated from the eyestalk neural tissue of the crab Oziotelphusa senex senex and used for cDNA synthesis. Total RNA of eyestalks was used earlier for the synthesis of cDNA in prawns Penaeus monodon (Davey et al., 2000; Udomkit et al., 2000, 2004; Krungkasem et al., 2002), Penaeus japonicus (Ohira et al., 2005) and Metapenaeus ensis (Gu et al., 2002), lobsters Homarus americanus (De Kleijn et al., 1998) and Nephrops norvegicus (Giulianini et al., 2002) and crab Charybdis feriatus (Chan et al., 1998).
In the present study, three CHH-family genes were cloned from the eyestalk neural tissue of the crab Oziotelphusa senex senex (Oss-SGP 1-3). Multiple genes of CHH-family peptides have been identified and cloned from several crustaceans. Tensen et al. (1991) have demonstrated the presence of two CHH genes in the eyestalks of American lobster Homarus americanus. Davey et al. (2000) have isolated five genes belongs to CHH-family hormones from the eyestalks of the prawn Penaeus monodon and named as SGP-I to SGP-V. In the same species, Udomkit et al. (2004) have cloned three CHH functional genes and designated as CHH-1, CHH-2 and CHH-3. In the freshwater prawn M. rosenbergii, two MIH/VIH-related peptide genes (Yang and Ranga Rao, 2001) and two CHH genes were cloned (Chen et al., 2003). Two genes encoding MIH isoforms were isolated from Fennropenaeus chinensis eyestalks (Wang et al., 2003). In the crab C. pagurus, Lu et al. (2000) have discovered three genes encoding MIH/MOIH. In American lobster Homarus americanus, Soyez et al. (1994) have discovered two genes encoding hyperglycemic hormone. In Procambarus clarkii and Procambarus bouvieri, only one CHH gene is present but hyperglycemic hormone is present in two isomorphs (Yasuda et al., 1994; Aguilar et al., 1995). These isomorphs are due to sterio-conversion of phenylalanine at 3rd position of mature hormone (Soyez et al., 1994).
In the present study, the amino acid sequences of partial mature sinus gland peptide(s) of the crab Oziotelphusa senex senex (Oss-SGP 1-3) were compared with the existing amino acid sequences of different crab species obtained from GenBank at NCBI using multiple sequence alignment. The three crab peptide sequences are showing 55.8-80.8% homology with each other. The obtained crab CHH-family peptides (Oss-SGP 1-3) exhibited less divergence with the CHHs of Pachygrapsus marmoratus (46.0%) and Scylla serrata (53.0%) and high divergence with the MIHs of Carcinus maenas, Cancer pagurus, Scylla serrata and Cancer magister (65.0-67.0%) and MOIH of Cancer pagurus (73.0%), indicating the obtained sequences in the present study are related to CHH-like peptides. Construction of full-length cDNAs for the sinus gland peptide gene(s) of Oziotelphusa senex senex is under progress in this laboratory, the results of which will throw more light in establishing the diversity of these genes.
In conclusion, we cloned three cDNAs encoding partial CHH proteins from Oziotelphusa senex senex neural tissue using PCR. By nucleotide and amino acid sequence comparison, crab neuropeptide cDNAs are closely related to the CHH of other crab species. Further study such as isolation of full length clones and determination of upstream elements may be needed to provide more information on the formation of isoforms of this gene in crab.
ACKNOWLEDGMENTS
We thank Department of Science and Technology, New Delhi for financial support in the form of research grant (SP/SO/C42/2001) to PSR. We also thank Mr. S. Umasankar for his help in maintaining the animals in the laboratory.