Research Article
A Semantic-Based Genetic Algorithm for Sub-Ontology Evolution
School of Computer and Information Engineering, Zhejiang Gongshang University, Xuezheng Street NO. 18, Hangzhou 310018, Zhejiang, Peoples Republic of China
Genetic Algorithm (GA) is a kind of efficient computational approach inspired from evolution theory and aimed to find optimum solution by searching problem space randomly. The theoretical foundations of GAs were first introduced by John Holland in early 1970s (Holland, 1975). Goldberg (1989) defined GAs as search algorithms based on the mechanics of natural selection and natural genetics. The GAs are used as a tool of optimization and searching in several brunches of science such as engineering, medical, business, statistics, signal processing etc. However, we face some difficulties when using canonical GAs mentioned above to solve optimization problems in semantic-based applications, because of the limitations or disadvantages of GA. In canonical GAs, chromosomes are often encoded as linear strings using binary or non-binary alphabet and genetic operators are performed based on probability and position. In the problem space of a semantic-based application, there are many complex semantic relations. We find that those methods cannot interpret the semantic relations in the problem space and hence overlook some useful information in evolution. Therefore, GAs are insufficient to satisfy the requirements in this case.
On the other hand, ontologies (Gruber, 1993; Van Heijst et al., 1997) provided a means of representing complex semantics in a formal and reusable from and play a critical role in building a large variety of semantic-based applications. The recent advent of the Semantic Web (Tim et al., 2001) has facilitated the incorporation of various large-scale on-line ontologies in different disciplines and application domains. For example, Unified Medical Language System (UMLS) (Bodenreider, 2004) included 975,354 concepts (a concept is a basic element denoting an object in ontology) and 2.6 million concept names in its source vocabularies for integrating biomedical terminology. Gene Ontology (Ashburner et al., 2000) included about 17,632 terms, 93.4% with definitions for gene product. Ontologies are increasingly seen as a key technology for enabling semantics-driven knowledge processing (Maedche et al., 2003; O’Leary, 1998).
Therefore, we try to use the formal semantics of ontology to improve canonical GA in several aspects and make it more adaptive to semantic-based problems. In this study, a new semantic-based GA is suggested in order to encode domain knowledge into the canonical GA for evolution based on semantics. Some advantages of this algorithm include the ability to express semantic information in chromosome representation and to preserve the information when applying genetic operators in evolution. We illustrated the feasible of the semantic-based genetic algorithm by applying it to solve the problem of sub-ontology evolution for dynamic ontology reuse.
Since, the emergence of GA, there have been various improved versions. Janikow (1993) argued that the two most important characteristics of GAs are robustness and the domain independence of their search mechanism. However, such an approach has both advantages and disadvantages. On the negative side lies the fact that the quality of the coding is crucial to the performance of GA. Operating in this space means using problem blind operators that often overlook some important information that could be utilized to guide the search. They propose a method of abstracting GA to the problem level for the supervised inductive learning. Initial results of their study indicate that GA can be effectively used to process high-level concepts and incorporate task-specific knowledge. Settea and Boullart (2000) introduced a Genetic-based Machine Learning application to achieve the automatic development of a rule set for an industrial production process. They argue that the basic representation scheme for chromosome was a simple string, which may in practice limit not only the class of suitable problems, but also the optimizing capabilities in the problem itself. Therefore, there is need for more complexity in the encoding structures of genetic mechanisms to escape from this rigid fixed string representation scheme. Kim and Cho (2005) argued that incorporating domain knowledge into evolutionary computation can improve the performance of evolved strategies and accelerate the speed of evolution by reducing the search space. They propose the systematic incorporation of opening domain knowledge and an endgame database into the framework of evolutionary checkers.
Compared with those improvements on GA, the focus of this study is to incorporate the formal semantics of ontology into GAs to solve semantic-based problems. The major difference with other methods is that semantics is used throughout the process of evolution in GA. We enhance the major components of canonical GA with formal domain knowledge.
PRELIMINARIES
In this study, we use Web Ontology Language (OWL) (McGuinness and Van Harmelen, 2004; Patel-Schneider et al., 2003) as the ontology language. Generally, an OWL Ontology (or simply an ontology), O is a pair <T, A>, where T and A are a T-box and an A-box, respectively in the Description Logic (DL) (Baader and Nutt, 2003). In OWL ontology, there is no direct definition about link; however, we can interpret property restrictions as cross-links between different classes.
Definition 1: (Triple): Given an ontology O = <T, A>, a triple, t, is represented as <c1, r, c2 >, where c1, c2 are concepts and c1, c2∈T and r denotes either a subsumption relation or a kind of cross-links.
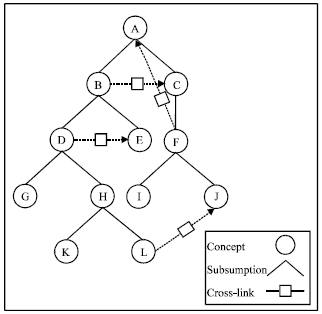 | |
| Fig. 1: | The graph representation for a sample ontology |
Therefore, an OWL ontology is treated as a set of semantically related triples. A triple refers to a set of statements in OWL ontology, abbreviated to <c1, r, c2>, which means that there is a relation between c1 and c2. Concepts as nodes and relations as arcs, an ontology can be represented as a graph. Figure 1 shows a sample ontology in form of graph. The number of arcs in the graph denotes how many triples there are in the ontology. Considering the direction of relation, the graph for an ontology is directed.
Definition 2: Relation path: Given a set of triples T = {ti| ti = <ai, ri, bi>, i = 1, 2, …, n} and two triples ts = <as, rs, bs>, tr = <ar, rr, br>, ts, tr ∈ T, if there exists m ≤ n - 2 and ts.bs ≡ ti.ai, ti.bi ≡ ti+1.ai+1, …, ti+m-1.b i+m-1 ≡ tr.ar, then we say there is a relation path from ts to tr.
Definition 3: Connective point: Given two triples t1 = <a, r, b> and t2 = <d, s, e>, if either a ≡ e or b ≡ d holds, then we say there is a connective point between t1 and t2.
A relation path in the ontology graph consists of a collection of triples, which connect to each other at connective points.
One of the most important tasks for ontology is reasoning. In OWL ontology setting, not all entailments are equally valued. Grau et al. (2006) shown three main reasoning tasks that ontology-based systems have traditionally focused on:
| • | Atomic concept satisfiability determines whether an atomic concept in the ontology is satisfiable |
| • | Classification computes the subsumption partial ordering of all the atomic concepts in the ontology |
| • | Instantiation and retrieval determine whether an individual is an instance of an atomic concept and retrieve all the instances of an atomic concept, respectively. |
Besides, we complement another reasoning task called indirect relationship, which determines whether or not there is a relation path between two concepts. For example, there is a relation path from I to A as we can see in Fig. 1, so I has an indirect relation with A.
Definition 4: Question: Given an ontology O = <T, A>, a question, q, is a triple <Qb, Qc, Qo>, where, Qo is an ontology, Qb is {ontology concept ci}, where ∀ci∈ Qo and Qc are a conditional string that denotes one of the reasoning tasks above.
We use question to represent a concrete ontology reasoning task.
SEMANTIC-BASED GENETIC ALGORITHM
We identify a canonical GA as these components: chromosome representation, fitness evaluation and genetic operators. We give semantic-based extension to each component in the following sub-sections. We present a semantic-based genetic algorithm (SemGA) to encode domain knowledge into GA and perform evolution based on semantics. We show the overall architecture of SemGA through Fig. 2. Compared with canonical GA, there is an additional semantic mediator as the mid-level between the components of GA and problems. The SemGA utilizes the semantics of domain ontology to solve semantic-based problems.
Chromosome representation: The first step in the implementation of any GA is to generate an initial population. In canonical GA each member of this population is a binary string of length L, which corresponds to the problem encoding. Each string is sometimes referred to as a genotype (Holland, 1975) or, a chromosome (Schaffer, 1987). In most cases the initial population is generated randomly. A chromosome is composed of a list of genes.
Chromosomes encoding in GA is to map the problem space to the parameter space. Possible solutions to the problem are represented as chromosomes by some encoding methods. We find that there are semantic relations between solutions to a semantic-based problem. We might overlook this information if we just use general linear string to represent chromosomes. Therefore, to a semantic-based problem, we use a composite structure for representing chromosomes, instead of linear string. If we represent the underlying semantics of a problem by a collection of triples, then a solution to the problem can also be described by a collection of triples.
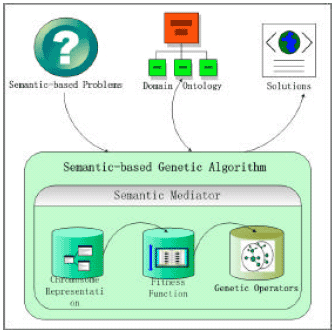 | |
| Fig. 2: | The abstract architecture of the semantic-based genetic algorithm |
Therefore, triple is used as the atomic component to express semantics of the problem space. We take all possible solutions to the problem as the problem space. Export all triples in the problem space as a triple list and each allele in a chromosome denotes a triple.
There are two possible methods to represent chromosomes semantically:
| • | Triple-based binary encoding: Export all triples in the problem space as a triple list. Each triple appearing in a solution will have the allele in the corresponding chromosome set to 1, or 0 otherwise |
| • | Triple-based non-binary encoding: A chromosome can be represented as a pair <E, T>, where E is the content of the solution and T = {triple ti}, a semantic description of the solution |
We show the two different encoding methods through the example in Fig. 3. Assume there are only three candidate solutions in the problem space. The chromosomes generated by the first method are shown in Fig. 3a and the second in Fig. 3b. As there is at most one role between two concepts, so role names for each triple are omitted in Fig. 3b. If a triple is represented as a binary character 0 or 1 by the triple-based binary encoding, we do not know whether or not two chromosomes have connective points in common, so it cannot preserve the connectivity of triples when we apply genetic operators like crossover to chromosomes.
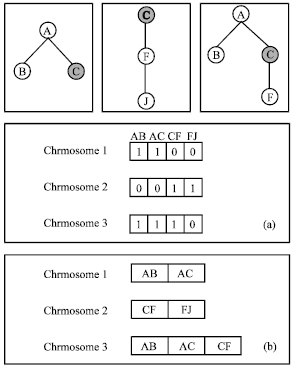 | |
| Fig. 3: | The results of the two different encoding methods to the same problem space |
In other words, the method overlooks some information in evolution. Therefore, we adopt the triple-based non-binary encoding in SemGA. We encode semantic entities including concepts and roles into chromosomes. A non-binary alphabet composed of concept names and role names in the problem space is used in chromosome representation. The length of chromosome is variable.
Fitness evaluation: The fitness evaluation function is a measure of performance for solutions. In order to determine an evaluation function for SemGA, we should think of the criteria for measuring whether a solution is better to the problem. As a chromosome for a solution is represented as a collection of triples in SemGA, we can evaluate the fitness value of a chromosome by computing the value of each triple. Let P be a population with n chromosomes. Assume the fitness evaluation function for triple is g(t), then the fitness value of a chromosome, S in P is calculated as follows:
| (1) |
where, gsum = ∑g(tj), tj is a triple in S, j = 1, 2, ..., m; Si ith chromosome in P
A chromosome with a collection of triples that have higher values gets a higher chance to survive in evolution. The overall fitness value of P is calculated as follows:
| (2) |
Genetic operators: After an initial population is generated, a GA carries out some genetic operators to generate offspring based on the initial population. Once a new generation is created, the genetic process is performed iteratively until an optimal result is found. We propose a collection of semantic-based genetic operators based on the chromosome representation and the fitness evaluation mentioned-before. Selection, crossover and mutation are three most important and common operators in SemGA.
Selection: When we apply the selection operator to a collection of chromosomes, we will compute the selection probability of each chromosome and chromosomes with large fitness values will gain a high chance to be selected into the next generation. The selection operator is almost the same as the one in canonical GA besides an additional comparing operation.
We will compare the semantic similarity between chromosomes. The semantic similarity is computed based on their Levenshtein distance (also known as edit distance) (Levenshtein, 1966). As the length of chromosome is not fixed, we use a relevant Levenshtein distance instead of the conventional one directly.
Definition 5: Relevant levenshtein distance: Given two sets S1, S2 and LD is the Levenshtein distance, then the relevant Levenshtein distance (RLD for short) between S1 and S2 is computed as follows:
| (3) |
Given two chromosomes S1 = <E1, T1>, S2 = <E2, T2>, the semantic similarity between S1 and S2 is equal to RLD(T1, T2).
Assume P is a population of chromosomes and the original selection operator is S (P). Then the semantic-based selection operator is described as follows:
| • | Compute the selection probability of each chromosome by its fitness value |
| • | Perform S(P) to get the candidate for the next generation |
| • | Compare the semantic similarity of chromosomes in P. If the semantic similarity of two chromosomes is arger than a threshold value, only one of them can be selected into next generation |
| • | Get the next generation P' |
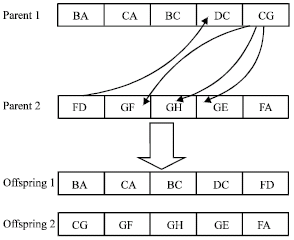 | |
| Fig. 4: | An example of the crossover operator |
Crossover: A single-point, triple-based crossover operator is applied, as shown in Fig. 4. We can apply the crossover operator to two chromosomes only when they are semantically related with each other, that is, there are connective points between them. Assume P is a population of chromosomes and the original crossover operator is C (Si, Sj), where, Si, Sj∈P. Then the semantic-based crossover operator is described as follows:
| • | Randomly select two different chromosomes from P, for say, S1 and S2 |
| • | Compare the semantic description of S1 and S2. Check whether or not there are connective points between them |
| • | If there exist connective points between S1 and S2, it means that they have some semantic relations. Therefore, it makes sense that we apply the crossover operator to the two chromosomes and get offspring. The crossover operator manipulates two chromosomes at the connective points where the triples in both chromosomes can be connected |
| • | Perform C (S1, S2) to get two offspring S1' and S2' |
The crossover operator can optimize the knowledge structure of parents. As the crossover operator manipulates two chromosomes only when there is connective point between them, it can optimize the semantic structure of parent chromosomes and make it more compact by generating offspring. Therefore, the overall structure of chromosomes is improved. For example, the two chromosomes in Fig. 4 have four connective points. We can randomly select a connective point, e.g., D between FD and DC, divide the two parents into four parts and then mate them to get two offspring.
 | |
| Fig. 5: | An example of the mutation operator |
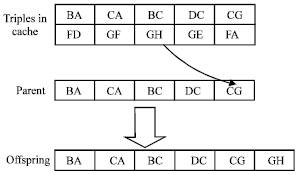
Mutation: A single-point, triple-based mutation operator is applied, as shown in Fig. 5. The operator adds one randomly chosen triple from the problem space to the chromosome or removes a triple from the chromosome. Assume P is a population of chromosomes and the original crossover operator is M (Si), where Si ∈ P. Then the semantic-based mutation operator is described as follows:
| • | Randomly select a chromosome from P, for say, S1 |
| • | Export all triples in the problem space as a triple list Lt |
| • | Compare the semantic description of S1 with Lt. Check whether or not there are connective points between them |
| • | Randomly select a triple from Lt and add the triple to S1; or randomly remove a triple from S1. If we want to add a triple to S1, we should check whether there is a connective point between the new-coming triple and the triples already in S1 |
| • | Perform M(S1) to get an offspring S1' |
The mutation operator enhances the chance for the evolution to jump out sub-optimization. Moreover, as the operator appends triple to a chromosome with some probability, it also enhances the chance for the chromosome to be connected with other chromosomes in the problem space. For example, to the chromosome in Fig. 5, we can randomly select a triple, GH from the triple list of the problem space and add it to the chromosome at a connective point, G to get an offspring.
SUB-ONTOLOGY FOR ONTOLOGY REUSE
A large-scale ontology like Gene Ontology contains relatively complete knowledge about the domain (e.g., gene product) it focuses on. The activities of many semantic-based applications rely only on localized information or knowledge (Ghidini and Giunchiglia, 2001). For example, a TCM ontology contains domain knowledge about TCM. If we want to build a semantic-based website about herbal medicine, we can extract a portion of knowledge about herbal medicine from the TCM ontology. All applications in that website are then implemented based on the portion of ontology, instead of accessing the complete TCM ontology. Our conjecture is that the activities of a specialized semantic-based application need only some specific aspects of a complete ontology. Small and context-specific portions of ontology are often needed in semantic-based applications especially those with limited capacity like embedded systems or mobile agents. This calls for the ability to extract from a large-scale ontology some context-specific portions and to allow them to evolve to achieve specialization. We represent and identify context-specific sub-ontologies (Mao et al., 2005) from the whole ontology and allow them to evolve according to past experience for gaining optimality.
Sub-ontology definition: In the following, we give a formal definition of sub-ontology to facilitate the discussion in the sequel.
Definition 6: (Sub-ontology): Given an ontology O = <T, A>, a sub-ontology (SubO for short), B, is a tuple <Bc, Bt, Ba, O>, where, Bt ⊆ T, Ba ⊆ A and C are {ontology concept ci}, where, ∀ci∈Bt. ∀ci and cj∈Bt, are is a path from ci to cj in Bt.
The concept set Bc denotes the context of a SubO and <Bt, Ba> denotes the local knowledge-base of the SubO. Given Bc, one can derive the knowledge-base of the SubO by searching the ontology O for the concepts in Bc. For example, to the sample ontology O in Fig. 1 and a concept set Bc = {B, A}, we can get the SubO in Fig. 6a. The SubO represents a indirect relation between the concept B and A.
Our definition of SubO is similar to the microtheory (Blair et al., 1992) in Cyc (Lenat, 1995). However, the difference is that microtheories in Cyc are static and invariant division of knowledge base, while SubOs can be extracted dynamically from source ontology and evolve in the process of semantic-based activities like reasoning (we will discuss the SubO evolution later). The evolution itself will not produce new knowledge, but it will improve the local knowledge structure of semantic-based systems, which can reuse SubOs in dynamic and adaptive way. Compared, with SubO, domain ontology can be treated as static and invariant resource to semantic-driven systems. Changes of domain ontology require owner privilege and domain experts intervention, so it is unpractical to let high-level systems directly modify and update source ontologies.
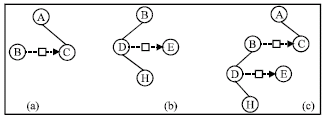 | |
| Fig. 6: | Several SubOs from the sample ontology |
However, different systems can have their local repositories of semantics from domain ontologies in terms of local and dynamic SubOs.
Ontology cache: As mentioned before, the context specific portions of ontology are represented as SubOs for semantic-based systems. Therefore, we can construct an ontology cache, which draws inspiration from the memory caching mechanism, to support SubO reuse. First emerging from data processing, caches work well because of a principle known as locality of reference. An ontology cache refers to a local repository for retrieved portions of ontology (in terms of SubOs) for future reuse. When a semantic-based system lacks the knowledge to perform reasoning tasks, it has to access ontologies. If the system extracts the required SubOs and keeps the most active ones in an ontology cache, the overall performance for reasoning is then optimized.
The domain knowledge involved in SubOs is reused to support reasoning tasks like a question. We try to answer questions by searching knowledge among a collection of SubOs in an ontology cache. For example, there is a question Q = <{H, B}, classification, Os>, where, Os refers to the sample ontology in Fig. 1. The meaning of Q is to check the subsumption between H and B. Assume there are three SubOs in an ontology cache (Fig. 6a-c). Then we find out SubO (b) in the cache and give a positive answer according to the semantics of SubO (b). When a collection of questions come, an ontology cache will try to answer each request by searching and reusing existing SubOs. If it can retrieve a SubO to answer the question, we say it is a cache hit, otherwise, a miss. If a cache miss occurs, the cache will try to extract SubOs from the source ontology. A newly-extracted SubO will be placed into the cache if the cache is not full; otherwise, the cache will try to replace the existing SubOs with the new one.
When a SubO is used to answer a question, not all parts in the SubO are equally used. When a triple is used as a part of proof for answering the question, it gets an incremental value.
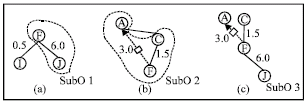 | |
| Fig. 7: | Optimizing SubOs in ontology cache |
Therefore, different triples in a SubO will gain different values. We define every m (m≥1) questions as a section. The cache value of a triple is updated every section. The initial value (section 0) of a triple is zero. The cache value of a triple, t is computed as follows:
| (4) |
| n | = | No. of section |
| λ | = | Expired rate of the cache value t got in the last section |
| Δcvn(t) | = | Hit number of t during section n |
Equation 4 takes into account both the past experience and recent performance for each triple. Then the cache value for a SubO, B, in section k is as follows:
| (5) |
where, n is the number of triples in B.
As the size of SubO is not fixed and there exists redundancy among SubOs, a SubO with a high value do not stand for a good SubO in an ontology cache. For example, there are two SubOs in an ontology cache in Fig. 7a-c. The cache value for SubO1 is (6.0+0.5)/2 = 3.25 and the cache value for SubO2 is (3.0+1.5+0.5)/3 = 1.67. It seems that SubO1 is a good SubO in the ontology cache. However, if we connect SubO1 and SubO2 at F and trim some triples with low cache value (less than 1.0), we can get a new SubO (Fig. 7c), the cache value of which is (3.0+1.5+6.0)/3 = 3.5. Therefore, we get a optimized SubO by connecting and trimming (a more complete evolution approach is discussed later). Therefore, we should go beyond using traditional cache replacement policy, in order to improve the performance of ontology cache. We must optimize the overall structure of SubOs in ontology cache. Such optimization can be achieved by some evolutionary computation algorithms like GA.
Therefore, the problem for SubO reuse in ontology cache can be summarized as follows:
| • | How to search knowledge contents in an ontology cache? |
| • | How to optimize the knowledge structure of SubOs in ontology cache for better performance? |
APPLY SEMGA TO SUB-ONTOLOGY EVOLUTION
The term Ontology Evolution in the area of ontology engineering is treated as a part of the ontology versioning mechanism (Klein and Fensel, 2001; Noy and Klein, 2004), which refers to access to data through different versions of an ontology. To contrast with ontology evolution, our concern of Sub-Ontology Evolution is inclined to evolve highly dynamic update of the SubOs in the local repository (ontology cache) of semantic-based system. We here adopt SemGA proposed earlier to solve the problem of structure optimization to support SubO caching.
Chromosome representation and fitness evaluation: We take the SubOs in ontology cache as the problem space. A SubO in ontology cache is represented a chromosome by the triple-based non-binary encoding in SemGA. We define two operators based on the chromosome representation for SubOs:
Encode: Given a SubO B = <Bc, Bt, Ba, O>, the encode operator proceeds as follows:
| • | Convert Bt into a set of triples T |
| • | Group <Bc, Ba, O> into a triple E |
| • | Return a chromosome S = <E, T> |
Decode: Given a chromosome S = <E, T>, where, E = <Bc, Ba, O>, the decode operator proceeds as follows:
| • | Convert T into an ontology T-box Bt |
| • | Return a SubO B = <Bc, Bt, Ba, O> |
The fitness function is a measure of performance for SubOs. In order to evaluate the fitness of chromosomes in evolution, we should think of the criteria for measuring whether a SubO is better to ontology reuse or not. The major objective of SubO evolution is to improve the cache performance. Therefore, the fitness function in our evolution approach mainly evaluates the fitness of a chromosome by the cache value of the corresponding SubO. A chromosome that has a higher cache value gets a higher chance to survive in evolution. Let P be a population with n chromosomes. We use Eq. 4 as the function g (t) in Eq. 1. Then the fitness value of a chromosome, S in P is calculated as follows:
| (6) |
| B | = | Corresponding SubO for S |
| Bi | = | Corresponding SubO for the ith chromosome in P |
According to definition 6, the concepts in a SubO should be connected. However, we may get disconnected SubO in evolution. In order to suppress the emergence of disconnected SubOs, we just give them a value of zero (fitness value should be non-negative). It makes sense that a connected SubO contains more information than disconnected one and thus should be preserved in evolution. For example, to the three SubOs in Fig. 6, SubO (c) has 5 triples, SubO (a) has 2 triples and SubO (b) has 3 triples. SubO (c) is as a combination of SubO (a) and SubO (b). There are 2 relation paths in SubO (a) and 5 in SubO (b). However, there are 12 paths in SubO (c). If one relation path is corresponding to a question, SubO (c) can answer more questions than SubO (a) plus SubO (b) together.
Genetic operators: SemGA operators are triple-based operations to optimize the semantic structure of SubOs in ontology cache. We implement the genetic operators of SemGA based on the SubO operators.
Selection: The selection operator uses the roulette wheel method, in which the selection probability of chromosome is defined below:
| (7) |
| n | = | No. of chromosomes in the population |
| f’(Si) | = | f (Si)/fmax |
| f(Si) | = | Fitness value of chromosome Si |
| fmax | = | Max (f(Sj)) |
| fsum | = | ∑f' (Sj) , j = 1, 2, ..., n |
The procedure of selection is as follows:
| Step 1: | Compute the selection probability Pi for chromosome i |
| Step 2: | Compute cumulative probability CPr = ∑Pi, i = 1, 2, ..., r |
| Step 3: | Let k = 1 |
| Step 4: | Generate a random number, η, uniformly distributed from the range 0.0 to 1.0. If CPi-1 < η ≤ CPi, select chromosome i as one of the elements in the next generation |
| Step 5: | Set k = k + 1 |
| Step 6: | If k = N (size of population), terminate; otherwise go to step 4 |
Crossover: Assume the population size is N and the crossover rate is Pc. The crossover operation is as follows:
| Step 1: | Let k = 1 |
| Step 2: | Randomly select two different chromosomes from the population. Randomly select a connective point between the chromosomes and divide each chromosomes into two set of triples at the point, one with the average cache value no less than the cache value of the whole SubO, the other with the average cache value less than the cache value of the whole SubO. Join the two parts with higher value from the two chromosomes together and the other parts together |
| Step 3: | Set k = k + 2 |
| Step 4: | If k is larger than N• Pc, stop; otherwise repeat step 2 |
The crossover operator can optimize the structure of parent SubOs and make it more compact. More important, the operator results in a better offspring and a worse offspring compared with the parents. The better offspring with high value will survive, while the other one will die. Therefore, the overall structure of ontology cache is improved.
Mutation: Assume the population size is N and the mutation rate is Pm. The mutation operation is as follows:
| Step 1: | Export all SubOs in the ontology cache as a triple list Lt |
| Step 2: | Let k = 1 |
| Step 3: | Select a chromosome from the population and generate a random integer, n, with a uniform distribution of (0, 1/Pm). If n equals to 0, a mutation is conducted to the chromosome. Randomly select a triple from Lt with high cache value and append it to the chromosome; or remove a triple with low cache value from the chromosome |
| Step 4: | Set k = k + 1 |
| Step 5: | If k is larger than N, stop; otherwise repeat step 3 |
Evolution procedure: We use SemGA to achieve SubO evolution based on the chromosome representation, fitness evaluation function as well as genetic operators before-mentioned. The evolution procedure is shown as follows:
| Step 1: | Extract SubOs from the source ontology according to some questions |
| Step 2: | Start evolution if the volume of the SubOs in the cache exceeds a predetermined level i.e., about 90% of the cache space is occupied by SubOs |
| Step 3: | Encode the SubOs in the cache as an initial population of chromosomes |
| Step 4: | After an initial population is generated, evolve the population based on SemGA. The genetic process is performed iteratively until an optimal result is found |
| Step 5: | Carry out some genetic operators to generate offspring based on the initial population |
| Step 6: | Once a new generation is created, compare the chromosomes in the population and merge ones with high similarity |
| Step 7: | Evaluate the fitness values of the chromosomes in the population. Terminate if the overall fitness is higher a threshold value; otherwise, go to step 5 |
| Step 8: | Decode the chromosomes in the result population to a set of SubOs and replace them with the original ones in the cache |
We can improve the performance of the SubO caching by using SemGA. The local knowledge structure of the ontology cache becomes more adaptive to answering questions via evolution.
EMPIRICAL RESULTS
Here, we examine the performance of SemGA by using a large-scale TCM ontology (Zhou et al., 2004). As the ontology is very large in scale, we only select several parts (Traditional Chinese Drug, Acupuncture, etc.) from the TCM ontology as the data-set for the experiment. There are about 3000 concepts in the selected ontology subset.
Experiment design: An important issue that impacts the performance of cache is the cache replacement policy. Such policies always apply a value function to each of the cached items and choose the items with the lowest values as victims. Therefore, we want to compare the SemGA approach with three traditional cache replacement policies: Random, FIFO and LRU. The experiment setting and major parameters are shown in Table 1. We have implemented four simple agents, each with an ontology cache and a cache replacement policy. With each agent, we perform the following procedure:
| Table 1: | The experiment setting and major parameters |
 | |
| Step 1: | Warm up the ontology cache by extracting SubOs from the source ontology according to some questions |
| Step 2: | Generate a collection of questions based on the TCM ontology |
| Step 3: | Submit the questions to the agent section by section. Questions are selected randomly. The agent answers questions by reusing existing SubOs in its ontology cache |
| Step 4: | For the agent with the SemGA policy, check the free space of its ontology cache. Perform evolution if the cache is nearly full. For other ontology caches, just skip this step |
The process of evolution is performed between the intervals of questions. This is to some extent similar with the behavior of prefetch, so we do not take into account the time of evolution when computing the cost of cache response.
| • | For each agent, calculate the average response time and hit-ratio of its ontology cache every section |
Compare cache performance: The primary metrics used to evaluate the performance of an ontology cache are hit-ratio and response time. A cache with higher hit-ratio and less response time is better.
The response time for ontology cache is as follows:
| (8) |
| Tpenalty | = | Textraction + Tstore |
| Tsearch | = | Time to search and retrieve a SubO from the ontology cache |
| Tanswer | = | Time to answer a question by reusing a SubO |
| Textraction | = | Time to extract a SubO from the source ontology |
| Tstore | = | Time to store a SubO into the ontology cache |
As the TCM ontology is stored in a relational database as a Jena (McBride, 2002) model and ontology cache is implemented in memory in this experiment, Tpenalty>> Tsearch and Tpenalty >> Tanswer. Therefore, we can ignore Tsearch and Tanswer when computing response time. The results presented here are a small, but representative set of the experiments we have run.
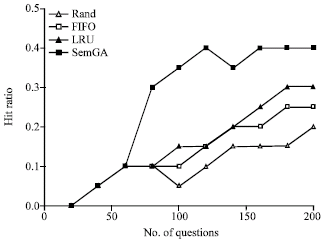 | |
| Fig. 8: | Hit-ratios for four cache policies |
The hit ratio for ontology cache is as follows:
| (9) |
| Nreuse | = | Number of reusing existing SubOs in the cache |
| ΔN | = | Section of questions answered within a time span |
Figure 8 shows the average hit-ratios for the four agents with different cache policies. The x-axis of the figure denotes the number of questions submitted to an ontology cache. It can be seen that four policies provide different performance across the range of questions. From the Fig. 8, we can see that SemGA does the best followed by LRU, FIFO and then Random. In the beginning, there is enough free space for SubOs in the SemGA cache, thus far from performing evolution. Therefore, the performance of the SemGA cache is fairly close to the LRU cache. After answering a batch of questions, the volume of the SubOs in the cache exceeds Le and the SemGA cache starts to evolve the SubOs. When evolution occurs (at the point of 60 questions), the performance of the ontology cache will be improved a lot. After several generations of evolution, the SemGA cache performs much better than the LRU cache. Figure 9 shows the average response time for the four agents with different cache policies. As response time is directly related with hit-ratio, the result in Fig. 9 is similar with that in Fig. 8 and the SemGA cache performs best in terms of response time. Generally speaking, the average response time for the SemGA cache is decreased by evolution.
Analyze fitness value: We now investigate the fitness values of chromosomes in evolution. To the SemGA cache, we record the fitness values of the chromosomes in every generation. Figure 10 shows a plot of the fitness values for the chromosomes after each generation of evolution, which is used to evaluate the overall fitness of the population.
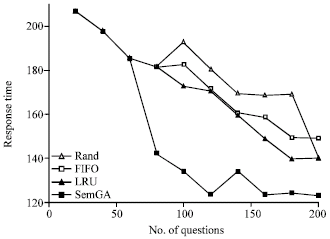 | |
| Fig. 9: | Response time for four cache policies |
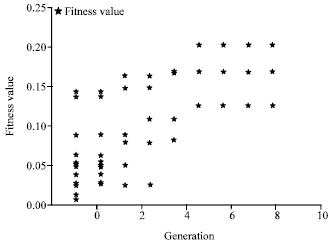 | |
| Fig. 10: | The fitness value of the chromosomes in evolution |
As can be seen in the Fig. 10, the fitness values of chromosomes are polarized to the two ends after several generations of evolution, a few of chromosomes with high values and a number of chromosomes with zero value. It is mainly because the crossover operator can generate a better offspring and a worse offspring compared with the parents. Although, the chromosomes with zero value increase, the chromosomes with high value also increase via evolution. As the chromosomes with zero value cannot pass the selection, the total number of chromosomes is reduced via evolution. The evolution produces a collection of chromosomes with high fitness values. It means the knowledge in the cache clusters into more compact and useful SubOs. Therefore, we get a new set of the SubOs with optimized knowledge structure and high cache values in the ontology cache.
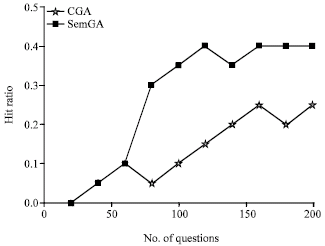 | |
| Fig. 11: | The hit ratios of CGA and SemGA |
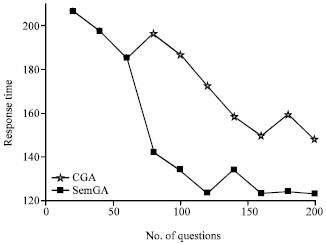 | |
| Fig. 12: | The response time of CGA and SemGA |
Besides, we find that the fitness values of the chromosomes become almost invariant after about 5 generations. The fitness values converge at the value of 0.16. It means the chromosomes have reached a stable states via evolution.
Compare with canonical GA: We also compare the performance of SemGA with canonical GA. We implement a canonical GA with the triple-based binary encoding and the genetic operators based on position and probability. As canonical GAs cannot ensure the connectivity of triples when applying genetic operators, we modify the fitness function for the canonical GA used in the experiment.
| (10) |
| B | = | Corresponding SubO for S |
| Bi | = | Corresponding SubO for the ith chromosome in P |
Compared with Eq. 6, it means we allow disconnected SubO in the canonical GA, which will not always get a value of zero in evolution.
We compare the performance of SemGA and canonical GA by following the same procedure for comparing SemGA with cache replacement policies. The results are shown in Fig. 11 and 12. As we can see, SemGA performs better than canonical GA in both response time and hit ratio to the problem of SubO evolution.
We present a semantic-based genetic algorithm called SemGA to incorporate domain knowledge into GA and perform evolution based on semantics. We identify a canonical GA as these components: chromosome representation, fitness evaluation and genetic operators and give semantic-based extension to each component. SemGA utilizes the formal semantics of domain ontology to solve semantic-based problems. We propose the concept of sub-ontology to represent context-specific portions from the whole ontology and reuse them in a cache-like repository. We identify the factor that impacts the performance of ontology cache is knowledge structure of SubOs. Therefore we propose to apply SemGA for SubO evolution to optimize the knowledge structure of ontology cache. We have designed a simulation experiment to evaluate the proposed approach with a large-scale TCM ontology.
There is still much room for improvement on the proposed algorithm. Future research issues include (1) improve the algorithm to get better speed and performance, (2) evaluate the algorithm with more complex and large-scale domain ontologies and (3) extend the evolution approach to a multi-agent environment and study the knowledge distribution in such an environment.
The study was partially supported by a Science and Technology Department of Zhejiang Province Program (NO. 2008C33G1770004) and a Science and Technology Program of ZJGSU (NO. 1130XJ200920).