ABSTRACT
Helicobacter pylori (HP) is the main cause of human chronic gastroduodenal disease including gastric cancer. Study of the protein interaction of the pathogen and its host will contribute to the understanding of the pathogenesis. The existing studies provide only static information of protein interaction and the lack of dynamics mechanism and system model of their protein interaction cannot explain the process of pathogenic and the diversity of clinical outcome. This study introduces system (network) biology to establish the dynamics model of the relation of HP virulence factors and its protein network, the structure and state model of host protein network under the conditions of different gastroduodenal diseases, the structure and dynamics of HP protein network exerting influence on its host protein network by way of coupling and the state transition model of the host protein network drived into a gastroduodenal disease status by HP protein network. Then, these models are systematized to be the model of the pathogenesis of gastroduodenal disease from which further discussions are also made.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/itj.2013.429.433
URL: https://scialert.net/abstract/?doi=itj.2013.429.433
INTRODUCTION
Helicobacter pylori (HP) is main cause of human gastric and duodenal disease (El-Omar et al., 2000). Epidemiological data show that more than half of the world population have HP infection (Mahdavi et al., 2002) but most of them do not occur stomach lesion, or mild chronic gastritis and only some develop atrophic gastritis, peptic ulcer disease, gastric cancer, gastric mucosa associated lymphoid tissue lymphoma (MALT ) or other serious diseases. Matteo et al. (2010) showed the diversity of HP infection clinical outcome is not only related to the different pathogenic of HP virulence factors, but also related to the interaction of HP and host and individual response. Most of serious diseases associated with HP occur in the circumstance of the coming together of specific virulence factor (such as CagA, VacA) of HP and susceptibility genetic background of the host. The lack of the information of the interaction of the pathogen and its host directly affects pathogenesis understanding. HP protein plays not only an important role in their pathogenicity and antigenicity (Zhang et al., 2006) but also causes gastrointestinal disease by way of the interaction with the host protein (Dubreuil et al., 2002). It is necessary to analyze the interaction of HP and its host based on the coupling of their protein networks (Li et al., 2011). Such research is being hotspot of the study of gastroduodenal pathogenesis (Zhang et al., 2011).
CURRENT STUDY OF HP-HOST PROTEIN INTERACTION
The first HP protein interaction map including 1200 proteins was drawn by Pasteur Institute in France by way of yeast two hybrid system (Rain et al., 2001). The interactions between 6559 of human protein and 623 of HP protein were predicted with the aid of a homology search method (Tyagi et al., 2009), including 549 HP coded proteins which their genome function not yet clearly known and 13 secreted proteins which experimentally confirmed and identified 833 interactions involved with the extracellular domain of HP transmembrane proteins. It was confirmed that (Jungblut et al., 2010), the type IV secretion system (T4SS) encoded by HP Cag pathogenicity island can make CagA protein transfer to gastric epithelial cells. For a TN2, there are other proteins involved in the functioning of CagA apart from the type IV secretion system (Baek et al., 2007). It is found that expression profiling of the host protein also has an obvious change in the infection. After invasion into AGS (gastric mucosa epithelial cells) (Das et al., 2005), HP 11637 CagA protein causes the host expressing αPix which interacting with CagA. The study of four kinds of gastric epithelial cell before and after the infection found biological marker protein Annexin II playing an important role in gastric cancer (Chan et al., 2006) and its expression quantity increasing in gastric epithelial cells after the infection, indicating that the proteins are also involved in HP pathogenesis. There are 28 proteins changing in gastric cancer AGS after the infection and among them 8 proteins is high expression in more than 60% samples of the tumor tissue (Backert et al., 2005). In the study of AGS infected by wild type and the same gene T4SS mutant strain (Clyne and Drumm, 2004), the cytoskeleton actins signaling molecules of the host may be the target of virulence factor. In the comparison of the toxicity mechanism of CagA+and CagA- (Yinghui, 2011), there was no protein of tyrosine phosphorylation found in a host infected by CagA+strain, while two proteins of tyrosine phosphorylation were detected in another infected by CagA-strain, prompting that the phosphorylation of CagA may inhibit phosphorylation reaction of other proteins. The process of HP cause gastritis will also make the balance of Th1/Th2 bias in the Th1 (Yan et al., 2011), resulting the production of a large number of type Th1 cytokine including key molecule such as IFN-γ (Interferon-γ) of Th1 response. In another study (Kim and Kim, 2009), immune protein induced by HP infection and oncogene protein are found interaction to each other.
Studies have shown that the relation of the pathogen and the host is very complex and can be attributed to the following aspects from protein perspective:
| • | HP virulence factors appear in some strains but do not show in others and a virulence factor may or may not occur in some periods of a same strain. Expressing or not of a virulence factor and expression intensity are related to several other proteins |
| • | In the presence of a HP toxic factor, some hosts are not manifesting disease and other are suffering from gastroduodenal diseases. A certain toxic factor may make different host getting different diseases, respectively such as chronic gastritis, gastric ulcer, duodenal ulcer or gastric cancer and so on and proteins interacting with toxic factors are also affected by other proteins inside the host |
| • | A causative factor may interact with a variety of host proteins and a host protein may also be associated with multiple pathogenic factors. A HP protein may also be at a number of pathogenic related signaling paths, so is a host protein |
Although, providing plenty of static information of protein interaction on the microcosmic as above, previous researches can not only not describe intricate relation of the pathogen and the host and the dynamic process of the pathogenic, but also not explain how the HP virulence factors interact with different hosts of different genetic backgrounds and how the host react to the infection according to its genetic characteristics so as to suffer different diseases.
MOTIVATION OF THE STUDY
The process of gastrointestinal disease induced by the pathogen is involved not only with the internal dynamics of the bacteria, but also with the host itself genetic characteristics and the external interaction between the two sides. Therefore, system (network) biology viewpoint and method are introduced to take protein interaction of HP and its host as a relatively whole system which is composed of three subnetworks (subsystems), such as a HP protein network, a host protein network and a coupled network coming from the interaction of the former two. Based on the study of transmission dynamics, synchronization dynamics of the subsystems, several models will be established, such as a dynamics model of the relation of HP virulence factors and its protein network, a structure and dynamics state model of the protein network of patients with gastrointestinal disease, a coupling structure and dynamics model of HP protein network dynamics state exerts influence on host protein network and a state transition model of host protein network driven by HP into the state of gastrointestinal disease. After the integration of above models, a gastroduodenal disease pathogenic model will be obtained, through which not only the pathogenic mechanism will be elucidated but correlation of host genetic characteristics and gastroduodenal disease also analyzed and diversity clinical outcome of HP infection also interpreted.
RESEARCH PROGRAMME
Topological structure of protein interaction networks: Based on a basic framework coming from HP protein-protein interaction maps established by the Pasteur Institute, an initial structure of HP protein network is to be formed according to BIND (Bimolecular Interaction Network Database), DIP (Database of Interacting Proteins) and other sources, so is that of the host protein network to be formed from the integration of BIND, DIP and other infection data according to a basic framework derived from HPRD (Human Protein Reference Database).
After that, false positive protein-protein interaction is removed according to semantic similarity of GO (Gene Ontology) phrase (Mahdavi and Lin, 2007) and complete and reliable topological structure of the protein networks of HP and the host are gotten, respectively. Based on GO phrase semantic, the method to calculate the similarity of two proteins is:
| (1) |
If SimP (PA,PB) greater than a certain threshold, the interaction of the two proteins is considered to be true, or else to be false. The experimental route is as Fig. 1.
Dynamics model of protein interaction networks: The transmission and synchronization dynamics of HP protein network is studied under the condition of CagA, VacA and other major virulence factors, so as to identify relationship between virulence factors and network dynamical attractors. Then, the characteristics of protein networks of abnormal hosts which suffer different diseases such as mild chronic gastritis, atrophic gastritis, peptic ulcer disease, gastric cancer, gastric mucosa associated lymphoid tissue lymphoma are analyzed and the protein network dynamics states of these diseases are found out. The node state of discrete model of protein network global dynamical properties is described as follows:
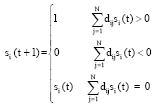 | (2) |
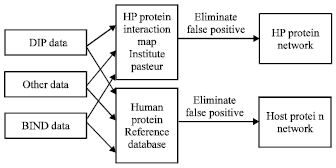 | |
| Fig. 1: | The experimental route for getting initial structures of protein networks of HP and the host |
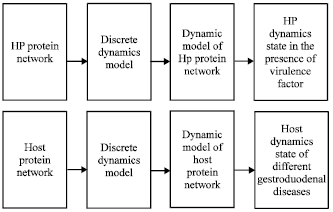 | |
| Fig. 2: | The experimental route for study of network dynamics |
Let S ε{X, Y}, S = X meaning node set of HP protein network while S = Y that of the host protein network. si (t+1) is the state of node i in time (t+1), while D = (dij)εRNxN is the interaction strength of protein j on i. The experimental route is shown in Fig. 2.
Structure and model of the coupling of the protein networks of hp and the host: HP and the host protein networks are both complex system (complex network), while each of them is only a subsystem (subnetwork) of a much more complex one. In this study, gastrointestinal disease is taken as the evolution result of a complex system composed from HP and the host and as the product of the dynamics process based on the coupling of the two protein networks. According to the node state given in formula (2) as above, the two networks are expressed as follows (Li et al., 2009):
| (3) |
| (4) |
where, xi and yi respectively are the nodes of HP and host protein networks when the s of (2) getting different values; f (x) and g (x) are the functions of nonlinear continuously differentiable; μ and ε, respectively are the connection strengths of internal nodes of the networks; A = (aij)εRNxN and B = (bij)εRNxN are internal connection matrixes of the networks, while P = (pij)εRNxN and Q = (qij)εRNxN are their external connection matrixes and ρ is their coupling strength. The topologies of the two protein networks are respectively imported into the formulas to get corresponding network models. Based on them, the signal transduction pathway and coupling dynamics are studied. The experimental route of this part is Fig. 3.
Driving and transfering models of the coupled networks: In the circumstances of CagA, VacA and other major virulence factors or their combinations, the mechanism of the propagation and synchronization taking place in the coupling of the two networks are analyzed and\ the dynamics process of gastrointestinal disease is studied.
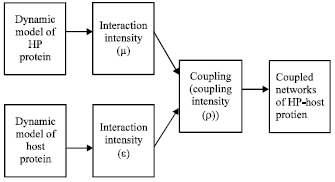 | |
| Fig. 3: | The experimental route for study of the coupling model |
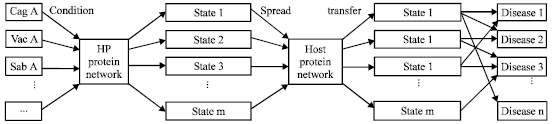 | |
| Fig. 4: | The experimental route for study of the driving and transfering of the coupled networks |
More over, the diversity of clinical outcome of the infection is to be further discussed and the correlation between the gen (Fig. 4). The experimental route for study of the driving and transfering of the coupled networks.
CONCLUSION
The study introduced system biology to establish several dynamics submodels and systematize them to be the model of the pathogenesis of gastroduodenal disease. The reuslting model is not only to formalize the complex relation of protein interaction of the pathogen and its host but also to describe HP pathogenic processes from dynamics. While elucidating the mechanism of a variety of clinical outcomes, the study has also provided a new systematic model, method and platform for further research of gastroduodenal disease.
ACKNOWLEDGMENT
This work is supported by Educational Commission of Guangxi Zhuang Autonomous Region of China. No. 200911MS196 and No. 201106LX505.
REFERENCES
- El-Omar, E.M., M. Carrington, W.H. Chow, K.E. McColl and J.H. Bream et al., 2000. Interleukin-1 polymorphisms associated with increased risk of gastric cancer. Nature, 404: 398-402.
PubMed - Zhang, J., S.G. Ding, L.J. Zhong, S.R. Lin, B. Yang, J.R. Peng and Y.X. Lou, 2006. Difference analysis on proteome of Helicobacter pylori in patients with peptic ulcer, gastritis and gastric cancer. Zhonghua Yi Xue Za Zhi, 89: 2690-2694.
PubMed - Dubreuil, J.D., G. Del Giudice and R. Rappuoli, 2002. Helicobacter pylori interactions with host serum and extracellular matrix proteins: Potential role in the infectious process. Microbiol. Mol. Biol. Rev., 66: 617-629.
CrossRef - Zhang, L., P. Wang, S.L. Wei and C.J. Liu, 2011. Advances in relationship between gastric disease and polymorphisms in both helicobacter pylori virulence factors and host genetics. Yi Chuan, 33: 558-566.
PubMed - Tyagi, N., O. Krishnadev and N. Srinivasan, 2009. Prediction of Protein-protein interactions between Helicobacter pylori and a human host. Mol. BioSyst., 5: 1630-1635.
PubMed - Jungblut, P.R., F. Schiele, U. Zimny-Arndt, R. Ackermann and M. Schmid et al., 2010. Helicobacter pylori proteomics by 2-DE/MS, 1-DE-LC/MS and functional data mining. Proteomics, 10: 182-193.
PubMed - Baek, H.Y., J.W. Lim and H. Kim, 2007. Interaction between the Helicobacter pylori CagA and alpha-Pix in gastric epithelial AGS cells. Ann. N.Y. Acad. Sci., 1096: 18-23.
PubMed - Das, S., J.C. Sierra, K.V. Soman, G. Suarez and A.A. Mohammad et al., 2005. Differential protein expression profiles of gastric epithelial cells following Helicobacter pylori infection using ProteinChips. J. Proteome Res., 4: 920-930.
PubMed - Backert, S., H. Gressmann, T. Kwok, U. Zimny-Arndt and W. Konig, 2005. Gene expression and protein profiling of AGS gastric epithelial cells upon infection with Helicobacter pylori. Proteomics, 5: 3902-3918.
PubMed - Kim, K.K. and H.B. Kim, 2009. Protein interaction network related to Helicobacter pylori infection response. World J. Gastroenterol., 36: 4518-4528.
PubMed - Li, C., C. Xu, W. Sun, J. Xu and J. Kurths, 2009. Outer synchronization of coupled Discrete-time networks. Chaos, Vol. 19.
CrossRefDirect Link








