ABSTRACT
In order to investigate the relationship between hybridization compatibility and genetic distances and to provide genetics evidence for obtaining bridge species for the introgression from S. lycopersicoides to L. esculentum, 48 materials including 5 genotypes from S. lycopersicoides and 43 genotypes from Lycopersicon (9 species) were used in the present study. Genetic distances among S. lycopersicoides and 9 species from Lycopersicon were estimated by morphological markers and Random Amplified Polymorphic DNA (RAPD) markers, respectively and the results showed that genetic distances ranged from 0.304 to 0.406, with the Morder being L. chilense, L. peruvianum, L. glandulosum, L. pimpinellifolium, L. chmielewskii, L. hirsutum, L. cheesmanii, L. esculentum, L. pennellii and L. parviflorum from near to distant. Furthermore, hybridization compatibility of L. chilense, L. esculentum and L. pennellii with S. lycopersicoides (as male parent) was also studied and the results showed that pollen germination and pollen tube elongation were different among hybrid combinations.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/pjbs.2006.1160.1168
URL: https://scialert.net/abstract/?doi=pjbs.2006.1160.1168
INTRODUCTION
Tomato (Lycopersicon esculentum Mill.) as a worldwide economy crop is planted from 45°(S) to 65°(N) (Shen and Xu, 1957). Because of ‘bottleneck’ effects of modern breeding and long cultivation domestication, genetic background of tomato is gradually becoming narrow and genetic diversity is becoming more and more deficient (Du et al., 1999). So long as breeders relied entirely on genetic variation in the European sources and their derivatives, just as Rick said, the progress in improving yields, disease resistance and other desiderata was slow upto 1940. Bohn and Tucker didn’t firstly use wild species as genetic source of desired characters, until they discovered strong resistance to Fusarium wilt in L. pimpinellifolium in 1940 (Rick, 1988). Thereafter, exploitation of wild sources accelerated. According to the statistics, so far genetic resources resistant to 42 main diseases of tomato were discovered in related wild species and more than half of them were transferred to tomato. The tolerance to drought, salinity, low temperature and high soluble solid contents in some wild species were also found (Rick, 1988; Wolf et al., 1986; Kamps et al., 1987; Chetelat et al., 1995a,b; Rick and Chetelat, 1995; Chetelat et al., 1997). However, any material with high resistance to CMV was not discovered in genetic resources of tomato yet, which made breeders of tomato puzzled.
S. lycopersicoides, as related wild species, is close to tomato in Solanum genus and is a valuable breeding material of tomato for high resistance to CMV and excellent tolerance to cold stress. It is also a unique species that can cross with L. esculentum in Solanum genus, which greatly attracted tomato breeders half a century ago (Rick, 1951). Since Rick firstly obtained F1LS of intergeneric hybrid, which was produced by L. esculentum (cv. Pearon) as female parent and S. lycopersicoides (No.30382) as male parent through embryo culture (Rick, 1951), breeders have mainly focused on how to obtain fertile intergeneric hybrid between S. lycopersicoides and L. esculentum during the last half century (Rick, 1986; Chetelat et al., 1989; Chetelat et al., 1998; Guri et al., 1991; Hossain et al., 1994; Matsumoto et al., 1997). Due to its high sterility of pollens (pollens vigor are 0.5%), the F1LS could not be directly backcrossed to L. esculentum (as female parent) and it could not be acted as female parent because of its unilateral incompatibility (Rick, 1951). Through nearly half a century’s endeavor, the fertile intergeneric hybrid between S. lycopersicoides and L. esculentum was recently obtained by both sexual and asexual methods although no real application in tomato improvement was yet reported so far (Chetelat et al., 1998; Matsumoto et al., 1997).
In this study, genetic distances among S. lycopersicoides and 9 species of Lycopersicon were respectively estimated by morphological markers and random amplified polymorphic DNAs (RAPD) markers and hybridization compatibility of S. lycopersicoides and 3 species of Lycopersicon that were different to S. lycopersicoides in genetic distance were also studied. We attempt to discover the relationship between hybridization compatibility and genetic distances and try to provide genetics evidence for introgression from S. lycopersicoides to L. esculentum by bridge species.
MATERIALS AND METHODS
Plant material: Forty-eight materials were used in this research, including 5 genotypes of S. lycopersicoides and 43 genotypes of Lycopersicon (9 species) (Table 1).
Morphological markers: Twenty-two traits were respectively investigated at some growth stages as follows (The qualitative traits were transformed into different number, while the quantitative traits were average value of ten measurement).
| X1 | : | Growth type, 0-infinity, 1-definity |
| X2 | : | Leaf color, 0-green, 1- pale-green |
| X3 | : | Leaf shape, 1-normal, 3-potato leaf-like, 5-bracken, 7-web like |
| X4 | : | Seedling stem color, 0-purple, 1-green |
| X5 | : | Stigma, 0-non-exserted, 1-exserted |
| X6 | : | Flower color, 0-yellow, 1-bright yellow |
| X7 | : | Torus, 0-no, 1-yes |
| X8 | : | Tip fertility of anthers, 0-sterile, 1-fertile |
| X9 | : | Inflorescence type, 0- mono-raceme, 1-multi-raceme |
| X10 | : | Growing habit, 0- trail, 1-erect |
| X11 | : | Number of carpel, 0-2 carpels, 1- not 2 carpels |
| X12 | : | Foliole number |
| X13 | : | Style length (cm) |
| X14 | : | Sepal length (cm) |
| X15 | : | Petal length (cm) |
| X16 | : | Anther length (cm) |
| X17 | : | Leaf length (cm) |
| X18 | : | Leaf width (cm) |
| X19 | : | Style shape, 0-straight, 1-curly |
| X20 | : | Anther shape, 0-not equal length, 1-equal length |
| X21 | : | Anther crack type, 0-side crack, 1-top crack |
| X22 | : | Anther color, 0-yellow, 1-white |
| Table 1: | Plant materials used in the research |
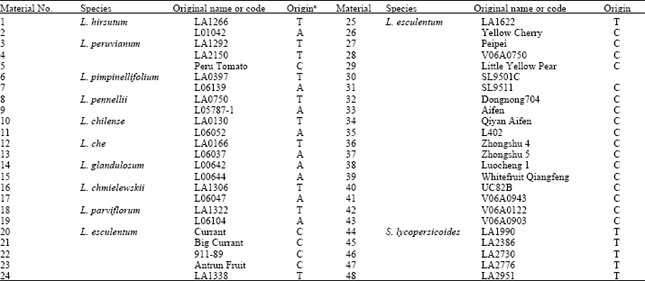 | |
A: Abbreviations of origin: T - C. M. Rick Tomato Genetic Resources Center, Dept. Vegetable Crops, University of California, Davis; A - Asian Vegetable Research and Development Center; C -China | |
RAPD markers: For each of 48 genotypes, young leaves from 10 plants were collected and total genomic DNA was extracted by the protocol of Zhao et al. (2002). The concentration and purity of exacted DNA were measured by spectrophotometry and the DNA concentration was adjusted to 20 ng μL-1. Using mixture of template DNA, 100 decamer oligonucleotide primers were screened, which came from OPA, OPB, OPC, OPD and OPN kit (20 primers in each Kit) purchased from Operon Technologies Inc. USA. According to stability and polymorphism of RAPD bands, 26 primers (OPA-03, 09, 11, 13, 14, 17, 18, 19; OPB-06, 07, 08; OPC-02, 05, 07, 09, 14, 15, 19; OPD-03, 13, 15, 18, 20 and OPN-03, 04, 13) were selected to amplify DNA in this research. The PCR volume was 25 μL containing 2 μL of adjusted genomic DNA, 2.5 μL 10 x PCR buffer (100 mM Tris-Cl buffer, 500 mM KCl, Gelatin 0.01%), 1.5 μL 25 mM MgCl2, 2 μL 2.5 mM dNTPs (Takara, Japan), 1 μL 10 μM primer, 0.2 μL (1 unit) Taq DNA polymerase (Takara, Japan) and 15.3 μL sterile water. Sterile mineral oil (20 μL) was added to each tube to seal the reaction mixture to prevent evaporation. PCR was carried out in a DNA Thermal Cycler (MJ PTC-100), with the following program: 94°C for 3 min followed by 35 cycles (94°C for 1 min, 36°C for 1 min, 72°C for 2 min) and finally at 72°C for 8 min. PCR products were electrophoresed on 1.4% (w/v) agarose gels containing ethidium bromide (EB) in 1 x TAE buffer. The gels were inspected and recorded under the UVP system (USA). Polymorphisms at all loci were confirmed by three repeated tests and each polymorphic RAPD fragment was scored 1 for presence and 0 for absence.
Data analysis
Morphological markers: Data of morphological markers were standardized using the following formulae:
![]() , i = 1, 2, 3, ……, n (sample numbers); j = 1, 2, 3, ……, m (variable numbers);
, i = 1, 2, 3, ……, n (sample numbers); j = 1, 2, 3, ……, m (variable numbers);
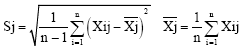 |
(Xij: the value of No.j feature of i sample, xij: the value of standardized Xij.
Sj: standard deviation of variable j; Xj: sample mean of variable j).
Maed software package was used to calculate genetic similarity matrix (Gs1ij) of 48 genotypes based on morphological markers and then dendrogram of cluster analysis was described according to the above genetic similarity matrix.
RAPD markers: Genetic similarity (Gs2ij) between every pair of 48 genotypes were respectively estimated according to Similarity for Qualitative Data (SIMQUAL) of NTSYS-pc package (1.80 version), using function of Sequential Agglomerative Hierarchical and Nested Clustering (SAHN) and cluster analysis was performed according to Unweighted Pair-group Method and arithmetic Average (UPGMA) linkage algorithm and dendrogram of cluster analysis was made by function of Tree Display.
Nei and Li method: Genetic similarity (Gs3ij) between each pair of 48 genotypes were estimated by RAPD markers, according to the formula proposed by Nei and Li (1979): Gsij = 2Nij/(Ni+Nj) (Nij: the number of the same amplified bands by sample i and j; Ni: the number of amplified bands by sample i; Nj: the number of amplified bands by sample j).
Average genetic distance: Average value of genetic similarity (Gsij) was calculated by the above three kinds of statistic methods, namely Gsij=(Gs1ij+Gs2ij+Gs3ij)/3 and corresponding average value of genetic distances of each pair were calculated as complements (1-Gsij) for Gsij.
Cross compatibility: L. esculentum (cv. White Fruit Qiangfeng), L. pennellii (LA0750) and L. chilense (L06052) were pollinated with the pollen of S. lycopersicoides (LA2386). Self-pollination of the above 4 parents was conducted in parallel as Control. After being pollinated of 2, 4, 8, 12, 16, 20, 24, 32, 40, 48, 60, 72 and 96 h, 5 flowers were respectively sampled from each treatment (including self pollination). All sampled styles with the ovary were dyed with aniline blue (Gao, 1981). The dyed tissue was identified under the Leica fluorescence microscope (Germany) for pollen germination and pollen tube elongation and photographed.
RESULTS
Morphological diversity: The 22 traits of Lycopersicon and S. lycopersicoides were investigated at certain growth stages. Based on the data of these traits, standardization was performed by Maed soft package and then genetic similarity coefficient of each pair of 48 genotypes was produced. Furthermore, genetic similarity matrix (Gs1ij) was produced (data not shown). The average genetic similarity of interspecies ranged from 0.9974 (S. lycopersicoides - S. lycopersicoides) to 0.1925 (L. parviflorum - L. pennellii).
The dendrogram of cluster analysis (data not shown) was described according to genetic similarity matrix and 48 genotypes were classified into 3 groups: A group contained all the species of Lycopersicon except L. pennellii, including 41 genotypes (i.e., 1-7 and 10-43, the serial number of genotypes as Table 1, hereinafter are same); B group contained all 5 genotypes of S. lycopersicoides (i.e., 44-48); C group is composed of 2 genotypes of L. pennellii (i.e., 8 and 9). Group A was still divided into 2 subgroups: A1 and A2. A1 was composed of 15 genotypes including most of wild species of Lycopersicon and a few semi-wild subspecies of L. esculentum (i.e., 1-7, 10, 11, 14-17, 20 and 24). A2 was mainly composed of cultivar sub-species and semi-wild sub-species of L. esculentum, including 21 genotypes (i.e., 19, 21-23, 25, 28-43). The rest 5 genotypes (i.e., 12, 13, 18, 26, 27) were not classified into A1 or A2.
RAPD diversity: Twenty-six decamer oligonucleotide primers were used in RAPD analysis for genomic DNA of total 48 genotypes. One hundred and forty-six of 201 detected loci were polymorphic, which occupied 72.64% of all the amplified loci. The results of RAPD were analyzed by NTSYS-pc. Genetic similarity coefficient among 48 genotypes tested ranged from 0.521 (LA0750-LA2730) to 0.959 (V06A0750-cv. Little Yellow Pear). Genetic similarity coefficient of 5 S. lycopersicoides genotypes and 43 Lycopersicon genotypes ranged from 0.521 (LA0750-LA2730) to 0.699 (V06A0122-LA2730) (Genetic Similarity matrix was not shown).
Clustering analysis was carried out by SAHN function and UPGMA method of NTSYS-pc software. Forty-eight genotypes of S. lycopersicoides and Lycopersicon were classified into 3 groups. Group A is composed of wild species of Lycopersicon including 15 genotypes (i.e., 1-15) and interestingly, among which genotypes 8 and 9 of L. pennellii were far away from the other 13 genotypes. Group B is composed of some wild species of Lycopersicon (i.e., 16, 18 and 19), 12 genotypes of semi-wild sub-species of L. esculentum (i.e., 20-31), the whole 12 genotypes of cultivar’s sub-species of L. esculentum (i.e., 32-43) and L06047 of L. chmielewskii (i.e., 17). Group C contains 5 genotypes of S. lycopersicoides (Fig. 1). Furthermore, group B could be still divided into 2 subgroups: B1 and B2. B1 contains 15 genotypes of some wild species of Lycopersicon and semi-wild subspecies of L. esculentum, while B2 contains the whole 12 genotypes of cultivar’s subspecies of L. esculentum L06047 was not classified into B1 or B2.
Genetic similarity matrix: Genetic similarity coefficients ranged from 0.736 (LA0750-LA2776) to 1 (all the genotypes to themselves) by Gsij=2Nij/(Ni+Nj) of Nei and Li (Data not shown).
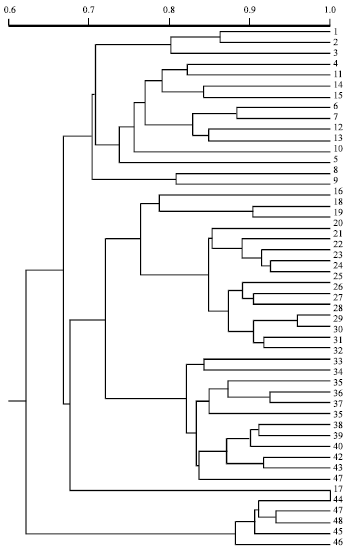 | |
| Fig. 1: | Dendrogram of RAPD cluster analysis using 48 genotypes of S. lycopersicoides and Lycopersicon (UPGMA) |
We also viewed genetic similarity coefficients of 5 genotypes of S. lycopersicoides and 43 genotypes of Lycopersicon with the values from 0.736 (LA0750-LA2776) to 0.843 (cv. V06A0122-LA2730). Moreover, interspecific average genetic similarity coefficients could be gained with the values from 0.755 (L. pennellii - S. lycopersicoides) to 0.969 (L. parviflorum -L. parviflorum).
Genetic distance between interspecies: The average genetic similarity matrix (Gsij) was produced by average value of corresponding genetic similarity coefficients of morphological markers (Gs1ij), RAPD markers (Gs2ij) and the third method of Nei and Li (Gs3ij), namely, Gsij=(Gs1ij+Gs2ij+Gs3ij)/3. The matrix of average genetic distances Dij could be also calculated as its complements (1-Gsij) (Nei and Li, 1979).
| Table 2: | Interspecific average genetic distance of S. lycopersicoides and Lycopersicon |
 | |
The matrix of interspecific genetic distance could be calculated according to Dij by average value of each species (Table 2). The maximum value is 0.498 (L. parviflorum - L. pennellii) and the minimum value is 0.032 (L. hirsutum - L. hirsutum). Genetic distances among S. lycopersicoides and each species of Lycopersicon ranged from 0.304 (L. chilense- S. lycopersicoides) to 0.406 (L. parviflorum - S. lycopersicoides), which ranked in the order of L. chilense, L. peruvianum, L. glandulosum, L. pimpinellifolium, L. chmielewskii, L. hirsutum, L. cheesmanii, L. esculentum, L. pennellii and L. parviflorum from near to far. This result was consistent with that of McClean and Hanson who adopted mtDNA in their research in 1986 (McClean and Hanson, 1986).
Cross compatibility: L06052 (L. chilense), White Fruit Qiangfeng (L. esculentum) and LA0750 (L. pennellii) as female parents, were respectively pollinated with the pollen of LA 2386 (S. lycopersicoides). Pollen germination and pollen tube elongation were observed under fluorescence microscope (using self-pollination as control).
Pollen germination: By self-pollination and interspecific hybridization, the pollen can germinate on the stigma normally, but the pollen germination of self-pollination after 2-5 h of pollination is obviously quicker than that of hybridization after 4-8 h. The germination of S. lycopersicoides’s pollen was so quick among the series of self-pollination that a lot of pollen of S. lycopersicoides germinated after 2 h of pollination. Among hybridization combinations, the pollen of S. lycopersicoides germinated most quickly on the stigma of L. chilense (1 - 7 in Fig. 2).
Elongation of pollen tube: Pollen tube of self-pollination elongated much more quickly than that of hybridization combinations. After 24 to 40 h of self-pollination, pollen tubes reached the bottom of the style or even entered the ovary (8-11 in Fig. 2). But quite a few of pollen tubes only reached the middle of the style after 4 days of pollination in hybridization combinations, among which pollen tube growth rates also had significant differences in different cross combinations. For example, some of pollen tubes had already extended to the bottom of the style or even entered the ovary after 48 to 72 h of pollination in the combinations L. chilense x S. lycopersicoides and L. esculentum x S. lycopersicoides, which pollen tube elongation of the former was a little faster than the latter (12, 13 , 15 and 16 in Fig. 2). While in the combination L. pennellii x S. lycopersicoides, pollen tubes still stayed in the middle of the style after 96 h of pollination, as well as after 30 h of pollination. In this case, pollen tube could hardly enter the ovary (14 in Fig. 2). To ensure the reliability of the results, sampling at some special time was repeated and the same results were obtained.
Furthermore, callose sediment was observed in the front of pollen tubes in the combinations L. pennellii x S. lycopersicoides and L. esculentum x S. lycopersicoides (14 and 15 in Fig. 2).
DISCUSSION
Genetic distance of S. lycopersicoides and Lycopersicon: Compared with other molecular markers, RAPD markers and associated technique (Welsh and McClelland, 1990; Williams et al., 1990) developed much more quickly, which have been widely used in many plant species for various research purposes (Hu and Quiros, 1991; Quiros et al., 1991; Demeke et al., 1992; Waugh et al., 1992; Foolad et al., 1993; Wilkie and Isaac, 1993; Jain et al., 1994; Mohan et al., 1994; Williamson et al., 1994; Paran et al., 1995).
Using RAPD markers, 48 genotypes in present study were classified into 3 groups, namely A (wild tomato species), B and C (5 genotypes of S. lycopersicoides), in which B could be divided further into 2 subgroups of B1 (a part of wild species and semi-wild subspecies of L. esculentum) and B2 (cultivated subspecies of L. esculentum). This result is basically consistent with the classification results using morphological makers in present research and with previous results (McClean and Hanson, 1986; Warnock, 1988; Lerfrancois et al., 1993).
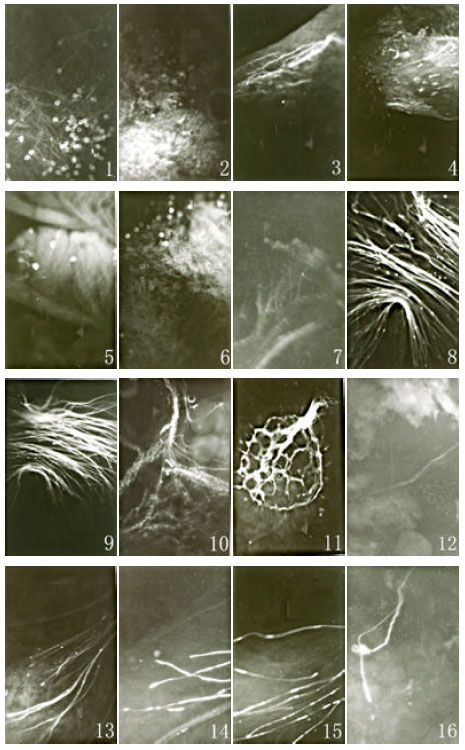 | |
| Fig. 2: | Pollen development (object lens x 5, ocular x 15). Nos. 1-7: Pollen burgeon at 4 h from LA2386 (No. 1), L. esculentum (No. 2), L. chilense (No. 3), L. pennellii (No. 4), L. esculentum x LA2386 (No. 5), L. chilense x LA2386 (No. 6) and L. pennellii x LA2386 (No. 7). Nos. 8-9: Pollen tube elongation of LA2386 (No.8, 24 h) and L. chilense (No. 9, 12 h). Nos. 10-12: Elongation of pollen tube to ovary of L. esculentum (No. 10, 24-32 h), L. chilense (No. 11, 24-32 h) and L. esculentum x LA2386 (No. 12, 60-72 h). Nos. 13-15: Pollen tube elongation of L. chilense x LA2386 (No. 13, 48 h), L. pennellii x LA2386 (No. 14, 96 h) and L. esculentum x LA2386 (No. 15, 48 h). No. 16: Elongation of pollen tube to ovary of L. chilense x LA2386 (60-72 h) |
Based on two kinds of makers (morphological and RAPD markers) in the present research, three kinds of statistical methods were used to calculate genetic similarity coefficients of each pair of 10 species including S. lycopersicoides and 9 species of Lycopersicon, by which genetic similarity would be much more reasonable and reliable. The results could provide some new information for studying the correlation between interspecies genetic distance and cross compatibility to find ‘bridge species’.
The significance of studying pre-fecundation’s compatibility between S. lycopersicoides and Lycopersicon and some approaches proposed to overcome the obstacles in distant hybridization: In the last half century, distant hybridization had become the main method to broaden genetic background of germplasm and to create new breeding materials (Chetelat et al., 1997; Rick, 1951; 1986; Chetelat et al., 1989; Chetelat et al., 1998; Menzel, 1962; Wann and Johnson, 1963; Xue, 1988; Xu et al., 1991; Liang and Mu, 1995; Li et al., 1998). For example, the resistances to Fusarium wilt and to Tobacco Mosaic Virus (TMV) in L. esculentum were from L. pimpinellifolium and L. peruvianum, respectively.
Owning to many barriers in distant hybridization, it was very difficult to transfer genetic traits from wild species to cultivars. By studying distant hybridization of Rhododendron genus, Williams and Knox (1982) proposed ten kinds of abnormalities such as twist, helix, swelling and callose plug of pollen tube on heterogenous stigma in 1982. In this research on the behaviors of pollen’s germination and pollen tube’s elongation of S. lycopersicoides on the stigma or in the style of Lycopersicon genus, we found that the percentage of pollen germination was low, elongation of pollen tube was slow and callose plug was in the front of pollen tube.
In the present research, L06052 (L. chilense), White Fruit Qiangfeng (L. esculentum) and LA0750 (L. pennellii) were pollinated with the pollen of S. lycopersicoides (LA 2386) and self pollination of 4 parents was conducted in parallel as control. The pollen could all be normally germinated (after 2-5 h of pollination) and pollen tubes reached the bottom of the style or even entered the ovary (after 24-40 h of pollination) in self-pollination. The results showed that pollen vigor of LA2386 and stigmas, styles and ovaries of other’s three female parents were normal.
The potentials of pollen germination and elongation of LA2386 on the stigmas and styles of others 3 female parents were distinctly deficient in comparison with self-pollination. The difference was also discovered among 3 hybrid combinations, which may be caused by difference of genotype. The pollen germination and pollen tubes elongation of LA2386 on the stigmas and styles of L. chilense were faster than those of L. esculentum and L. pennellii Those pollen tubes entered the ovaries in the L. chilense and L. esculentum, respectively for 60 and 72 h after pollination. While in L. pennellii pollen tubes only reached the middle of the style at 96 h after pollination. We have also measured that genetic distances of L. chilense, L. esculentum and L. pennellii are, respectively 0.304, 0.365 and 0.374 to S. lycopersicoides. It indicated that cross compatibility and genetic distances was relative. Therefore, it is possible to find ‘bridge species’ according to relationship between S. lycopersicoides and each species of Lycopersicon for introgression from S. lycopersicoides to L. esculentum.
During the process of the study on distant hybridization between S. lycopersicoides and Lycopersicon, we thought that pollen vigor of S. lycopersicoides was also an important factor (S. lycopersicoides pollen storage and flowering stage adjustment of Lycopersicon and S. lycopersicoides will be discussed in other report). Pollen vigor of S. lycopersicoides could maintain about 94.5% on the flowering day. From then on, it declined dramatically. On the 4th day after flowering, only 1.44% kept vigor. Sometime, distant hybridization could cause delayed pollen germination and slow elongation of pollen tube, resulting in the loss of sperm’s vigor, especially after more than 4 days of flowering.
Hybrid embryo rescue(Rick, 1951; Wu and Liang, 1992), in vitro fecundation and zygote culture are widely used to overcome barriers of distant hybridization (Smith, 1944; Thomas and Pratt, 1981). Pre-fecundation incompatibility can be decreased by chemical treatment and style shortening. However these methods are not stable and the ratio of success is too low. Although being a robust tool to overcome negative effects of maternal tissues and metabolites on pollen germination and pollen tube elongation, in vitro fecundation needs exquisite manipulation and necessary facilities (Yang and Zhou, 1998). Therefore, obtaining ‘bridge species’ of S. lycopersicoides and L. esculentum may be an effective approach for overcoming obstacle of distant hybridization.
Besides the above methods, genetic engineering is also a very efficient way for overcoming obstacle of distant hybridization. In our laboratory, some genes of interest (such as cf and Ve) from S. lycopersicoides have been cloned recently by cDNA library screening and rapid amplification of cDNA ends (RACE), which would be beneficial to improve genetic resource of tomato. Genetic engineering is possibly one of the effective approaches to achieve introgression from S. lycopersicoides to L. esculentum.
ACKNOWLEDGMENTS
We thank Dr. Chetelat (TGRC) and Mr. Yongguang Huang (AVRDC) for their kind provision of seeds of Solanum lycopersicoides and wild species of tomato. This research was funded by China Ministry of Education and Shanghai Agriculture Committee.
REFERENCES
- Chetelat, R.T., J.W.D. Verna and A.B. Bennett, 1995. Introgression into tomato (Lycopersicon esculentum) of the L. chmielewskii sucrose accumulator gene (sucr) controlling fruit sugar composition. Theor. Applied Genet., 91: 327-333.
Direct Link - Chetelat, R.T., J.W.D. Verna and A.B. Bennett, 1995. Effects of the Lycopersicon chmielewskii sucrose accumulator gene (sucr) on fruit yield and quality parameters following introgression into tomato. Theor. Applied Genet., 91: 334-339.
Direct Link - Chetelat, R.T., P. Cisneros, L. Stamova and C.M. Rick, 1997. A male fertile lycopersicon esculentum x Solanum lycopersicoides hybrid enables direct backcrossing to tomato at the diploid level. Euphytica, 95: 99-108.
Direct Link - Demeke, T., R.P. Adums and R. Chibbar, 1992. Potential taxonomic use of RAPD a case study in Brassica. Theor. Applied Genet., 84: 990-994.
CrossRef - Foolad, M.R., R.A. Jones and R.L. Rodriguez, 1993. RAPD markers for constructing intraspecific tomato genetic maps. Plant Cell Rep., 12: 293-297.
CrossRefDirect Link - Guri, A., L.J. Dunbar and K.C. Sink, 1991. Somatic hybridization between selected Lycoperscon and Solanum species. Plant Cell Rep., 10: 76-80.
Direct Link - Hu, J.G. and C.F. Quiros, 1991. Identification of broccoli and cauliflower cultivars with RAPD markers. Plant Cell Rep., 10: 505-511.
CrossRef - Kamps, T.L., T.G. Iskeib, R.C. Herner and K.C. Sink, 1987. Evaluation of techniques to measure chilling injury in tomato. HortScience, 22: 1309-1312.
Direct Link - Lerfrancois, C., Y. Chupeau and J.P. Bourgin, 1993. Sexual and somatic hybridization in genus Lycopersicon. Theor. Applied Genet., 86: 533-546.
Direct Link - McClean, P.E. and M.R. Hanson, 1986. Mitochondrial DNA sequence divergence among Lycopersicon and related Solanum species. Genetics, 112: 649-667.
Direct Link - Mohan, M., S. Nair, J.S. Bentur, U.P. Rao and J. Bennett, 1994. RFLP and RAPD mapping of the rice Gm2 gene that confers resistance to biotype 1 of gall midge (Orseolia oryzae). Theor. Applied Genet., 87: 782-788.
Direct Link - Quiros, C.F., J. Hu, P. This, A.M. Chevre and M. Delseny, 1991. Development and chromosomal localization of genome specific markers by polymerase chain reaction in Brassica. Theor. Applied Genet., 82: 627-632.
CrossRef - Rick, C.M., 1988. Tomato like nightshades affinities autoecology and breeders opportunities. Econ. Bot., 42: 145-154.
CrossRef - Warnock, S.J., 1988. A review of taxonomy and phylogeny of the genus Lycopersicon. Sci. Hortic., 23: 669-673.
Direct Link - Waugh, R., E. Baird and W. Powell, 1992. The use of RAPD markers for the detection of gene introgression in potato. Plant Cell Rep., 11: 466-469.
CrossRef - Welsh, J. and M. McClelland, 1990. Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Res., 18: 7213-7218.
CrossRefPubMedDirect Link - Williams, J.G.K., A.R. Kubelik, K.J. Livak, J.A. Rafalski and S.V. Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res., 18: 6531-6535.
CrossRefPubMedDirect Link - Williamson, V.M., J.Y. Ho, F.F. Wu, N. Miller and I. Kaloshian, 1994. A PCR-based marker tightly linked to the nematode resistance gene, Mi, in tomato. Theoret. Applied Genet., 87: 757-763.
CrossRefDirect Link - Wolf, S., D. Yakir, M.A. Stevens and J. Rudich, 1986. Cold temperature tolerance of wild tomato species. J. Am. Soc. Hortic. Sci., 111: 960-964.
Direct Link - Jain, A., S. Bhatia, S.S. Banga, S. Prakash and M. Lakshmikumaran, 1994. Potential use of random amplified polymorphic DNA (RAPD) technique to study the genetic diversity in Indian mustard (Brassica juncea) and its relationship to heterosis. Theor. Applied Genet., 88: 116-122.
CrossRefDirect Link








