Short Communication
Wolbachia Infection and Mitochondrial DNA Comparisons among Culex Mosquitoes in South West Iran
Department of Medical Entomology, School of Health, Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran
More than half of the population of the world lives in regions where mosquito borne diseases are widespread (TDR, 2009). The interruption of life cycle of the vector is the most effective method to control arthropod borne diseases. The extensive use of chemical insecticides against the vectors of diseases and harmful pests has resulted in the spread of insecticide resistance and relapse and recrudescence of these diseases (Gubler, 1998, 2009). Therefore, the investigation on the alternative methods which are safe in the environment is on the top of priorities.
Wolbachia is a group of bacteria belonged to alphaproteobacteria that live in the cytoplasm and transmit maternally to offspring of their hosts (O’Neill et al., 1992). About 66% of insect species are estimated to be Wolbachia infected (Hilgenboecker et al., 2008). These microbes manipulate the reproductive system of their hosts to increase their transmission probability within the infected population. Cytoplasmic incompatibility is the most widespread of these manipulation phenotypes of Wolbachia (Sinkins, 2004). On the basis of the above reason Wolbachia can be used as an important system to drive a suitable gene into the disease vectors population to disable them to transmit the disease agents, life-shortening or they can strongly inhibit the development of pathogens within the mosquito (McMeniman et al., 2009; Kambris et al., 2009, 2010; Moreira et al., 2009).
The biological control of mosquito in any area requires knowledge of field parameters such as the vector species, the parasites or related micro-organisms, the rate of natural infection and the interaction of the vector with the environment. Here field-collected Culex mosquitoes were examined for Wolbachia and in addition the diversity of mitochondrial DNA was examined in relation to Wolbachia infection.
Culex specimens were collected from Shoushtar in Khouzestan province in South West of Iran during April 2001. The Culex quinquefasciatus and Culex theileri samples were caught by hand aspirator during the day time from two different places in one village (Abeed haji baba) and Culex tritaeniorynchus were caught by CDC light trap during the night from one animal house of another village (Shoaebiae). The samples were then killed with chloroform and individually preserved dry in 1.5 mL microcentrifuge tubes under silica gel until identification in the laboratory. The samples were identified to species using morphological key of Zaim and Cranston (1986).
Total individual DNA was extracted from each single sample of Culex quinquefasciatus, C. tritaeniorynchus and C. theileri by the Livak DNA extraction procedure (Collins et al., 1987). They were examined for Wolbachia infection with PCR using the primers wsp. 81F (TGGTCCAATAAGTGATGAAGAAAC) and wsp. 691R (AAAAATTAAACGCTACTCCA) which were designed previously by Braig et al. (1998). The PCR conditions were as follows: 1X reaction buffer; 0.2 mM dNTPs mix; 5 mM MgCl2; 0.2 μM forward primer; 0.2 μM reverse primer; 0.04 u μL-1 Taq DNA polymerase; nuclease-free water and template DNA. PCR was performed in a 100 μL reaction mixture. The first seven reaction components were combined in a 1.5 mL microcentrifuge tube and were mixed gently. Then, 4 μL DNA template was added in 96 μL of reaction mixture in 0.2 mL thin walled PCR tubes, these were mixed and transferred to the PCR cycler with the following conditions: 94°C, 5 min (1 cycle); 95°C, 30 sec; 48°C, 1 min; 62°C, 2.5 min (40 cycles) and 72°C for 5 min (1 cycle). The PCR reaction products were analyzed by agarose gel electrophoresis of a 5 μL aliquot from the total reaction. The products were observed by UV transillumination of the ethidium bromide-stained gel.
The A+T rich region in Culex spp. was amplified using general primer SR-J-14612 (Simon et al., 1994) as forward and CxAT3 (Guillemaud et al., 1997) as reverse primer. Specific new primers were designed to amplify this region in C. tritaeniorynchus, C. quinquefasciatus and C. theileri. The new designed primers as indicated in Table 1 were as TR-ATF and TR- ATR to amplify an A+T region in C. tritaeniorynchus, QU,TH-ATF, QU-ATR and TH-ATR to amplify the region in C. quinquefasciatus and C. theileri.
| Table 1: | Primers used to amplify A+T rich region sequence and the percent of GC content |
 | |
RESULTS AND DISCUSION
Total 98 Culex spp. samples were collected in which 44 samples were identified as C. quinquefasciatus, 34 C. tritaeniorynchus and 20 C. theileri. All of the C. quinquefasciatus were infected with Wolbachia, based on PCR amplification of the wsp gene, while none of the C. tritaeniorynchus and C. theileri samples produced amplification of the Wolbachia wsp gene (Table 2).
The 340 bp of AT rich of mtDNA was sequenced from 30 individuals, 10 each of C. quinquefasciatus, C. tritaeniorynchus and C. theileri. The base composition of the mtDNA sequences as indicated in Table 3 is highly AT-biased with average of 89.1, 91.1 and 89.5% AT in the sequences of C. quinquefasciatus, C. tritaeniorynchus and C. theileri, respectively. Three sequence haplotypes were found in C. tritaeniorynchus and C. theileri while there was only one haplotype in C. quinquefasciatus.
Epidemics of the vector borne diseases such as malaria and arboviruses or re-emergence of them, following the economic and social changes after initial success, mean that these diseases still remains a major problem in many tropical or subtropical countries (Raghavendra et al., 2011; Gubler, 2009). The phenomenon of the cytoplasmic incompatibility of Wolbachia leading to spread in mosquito populations and its ability to induce inhibition of the development of viruses and parasites in the vector, plus the effects of lifespan-shortening this micro-organism made it a suitable way to prospect to control the vector borne diseases and malaria (McMeniman et al., 2009; Moreira et al., 2009; Kambris et al., 2009, 2010).
Study of Wolbachia infection in the field is an important step to evaluate the Wolbachia as a driving gene system (Turelli and Hoffmann, 1999). Wolbachia frequency and density in hosts vary significantly in nature compared to laboratory strains of the same species.
| Table 2: | The number of mosquito individuals, Wolbachia infection and the percent of Wolbachia infection in each species |
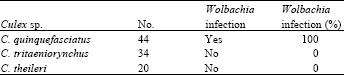 | |
| Table 3: | The number of mosquito individuals, sequence size (bp), the percent of AT, the number of mitochondrial haplotypes within each group of Culex species based on the AT-rich region sequence and the positions of nucleotide changes |
 | |
Part of these variation it is not only dependent on the genome of the host but environmental factors such as temperature can play a very important role in the varieties (Echaubard et al., 2010). In the collection region the temperature reaches more than 50°C in summer time and since high temperatures are known to be a cause of Wolbachia loss in laboratory studies, this could in theory reduce the incidence of Wolbachia. However, all 44 specimens of C. quinquefasciatus examined were positive for Wolbachia.
Wolbachia infection was not seen in any C. tritaeniorynchus samples and these negative results were similar with another study of Wolbachia incidence in Culex spp. in Thailand (Tiawsirisup et al., 2008). Also, positive amplification of the Wolbachia gene was not seen in C. theileri, a species which is considered as the main vector of Dirofilaria immitis (Nematoda: Filarioidea) on Madeira Island, Portugal (Santa-Ana et al., 2006). No previous studies have examined Wolbachia presence or absence in this species.
In the current study, the AT region as part of the mtDNA genome was sequenced from individuals of the Culex species C. quinquefasciatus, C. tritaeniorynchus and C. theileri. Haplotype diversity was just one for the Wolbachia infected C. quinquefasciatus while three haplotypes were seen in the uninfected populations (C. tritaeniorynchus and C. theileri). This reduced mtDNA diversity seen in C. quinquefasciatus may have been the result of the spread of Wolbachia on a relatively recent evolutionary time scale (since both Wolbachia and Mitochondria are solely maternally inherited, there is complete linkage between them and thus spread of a mtDNA variant by hitchhiking is associated with the spread of a Wolbachia infection. The presented results are supported by previous studies of mitochondrial genes and Wolbachia infection (Turelli et al., 1992; Ballard and Kreitman, 1994; Rand et al., 1994; Ballard et al., 1996; Johnstone and Hurst, 1996; Rigaud et al., 1999; Shoemaker et al., 1999; Ballard, 2000). Low intraspecific mtDNA variation can also result from some demographic event affecting females such as recent bottlenecks, low effective population size or from a recent selective sweep of a favoured mtDNA variant (Johnstone and Hurst, 1996), further field and population genetic studies could be used to clarify these issues.
Our data, for the first time, showed that Culex theileri samples were negative for Wolbachia. While, the Culex quinquefasciatus samples were positive with only one haplotype of mitochondrial genome. Regarding these field data, conducting a local genetic control programme and laboratory experiments, to horizontal transfer of Wolbachia strains to main vectors of important vector borne disease will be considered.
I would like to thank Dr. Steven Sinkins for his invaluable comments on the manuscript and I thank The Ministry of Health and Medical Education of Iran and Liverpool School of Tropical Medicine for their help.